+Search query
-Structure paper
| Title | W546 stacking disruption traps the human porphyrin transporter ABCB6 in an outward-facing transient state. |
|---|---|
| Journal, issue, pages | Commun Biol, Vol. 6, Issue 1, Page 960, Year 2023 |
| Publish date | Sep 21, 2023 |
 Authors Authors | Sang Soo Lee / Jun Gyou Park / Eunhong Jang / Seung Hun Choi / Subin Kim / Ji Won Kim / Mi Sun Jin /  |
| PubMed Abstract | Human ATP-binding cassette transporter subfamily B6 (ABCB6) is a mitochondrial ATP-driven pump that translocates porphyrins from the cytoplasm into mitochondria for heme biosynthesis. Within the ...Human ATP-binding cassette transporter subfamily B6 (ABCB6) is a mitochondrial ATP-driven pump that translocates porphyrins from the cytoplasm into mitochondria for heme biosynthesis. Within the transport pathway, a conserved aromatic residue W546 located in each monomer plays a pivotal role in stabilizing the occluded conformation via π-stacking interactions. Herein, we employed cryo-electron microscopy to investigate the structural consequences of a single W546A mutation in ABCB6, both in detergent micelles and nanodiscs. The results demonstrate that the W546A mutation alters the conformational dynamics of detergent-purified ABCB6, leading to entrapment of the transporter in an outward-facing transient state. However, in the nanodisc system, we observed a direct interaction between the transporter and a phospholipid molecule that compensates for the absence of the W546 residue, thereby facilitating the normal conformational transition of the transporter toward the occluded state following ATP hydrolysis. The findings also reveal that adoption of the outward-facing conformation causes charge repulsion between ABCB6 and the bound substrate, and rearrangement of key interacting residues at the substrate-binding site. Consequently, the affinity for the substrate is significantly reduced, facilitating its release from the transporter. |
 External links External links |  Commun Biol / Commun Biol /  PubMed:37735522 / PubMed:37735522 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 3.9 Å |
| Structure data | EMDB-36937, PDB-8k7b: EMDB-36938, PDB-8k7c: |
| Chemicals |  ChemComp-PEV: 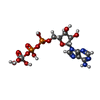 ChemComp-AOV:  ChemComp-MG:  ChemComp-ADP: 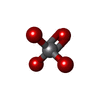 ChemComp-VO4: |
| Source |
|
 Keywords Keywords | TRANSPORT PROTEIN / ATP-Binding Cassette transporter mitochondrial transporter / post-occluded / ADP/VO4-bound / ABCB6 / MEMBRANE PROTEIN / ATP-Binding Cassette transporter / mitochondrial transporter / outward-facing state |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers







 homo sapiens (human)
homo sapiens (human)