+Search query
-Structure paper
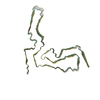
| Title | Cryo-EM structure of anchorless RML prion reveals variations in shared motifs between distinct strains. |
|---|---|
| Journal, issue, pages | Nat Commun, Vol. 13, Issue 1, Page 4005, Year 2022 |
| Publish date | Jul 13, 2022 |
 Authors Authors | Forrest Hoyt / Heidi G Standke / Efrosini Artikis / Cindi L Schwartz / Bryan Hansen / Kunpeng Li / Andrew G Hughson / Matteo Manca / Olivia R Thomas / Gregory J Raymond / Brent Race / Gerald S Baron / Byron Caughey / Allison Kraus /  |
| PubMed Abstract | Little is known about the structural basis of prion strains. Here we provide a high (3.0 Å) resolution cryo-electron microscopy-based structure of infectious brain-derived fibrils of the mouse ...Little is known about the structural basis of prion strains. Here we provide a high (3.0 Å) resolution cryo-electron microscopy-based structure of infectious brain-derived fibrils of the mouse anchorless RML scrapie strain which, like the recently determined hamster 263K strain, has a parallel in-register β-sheet-based core. Several structural motifs are shared between these ex vivo prion strains, including an amino-proximal steric zipper and three β-arches. However, detailed comparisons reveal variations in these shared structural topologies and other features. Unlike 263K and wildtype RML prions, the anchorless RML prions lack glycophosphatidylinositol anchors and are severely deficient in N-linked glycans. Nonetheless, the similarity of our anchorless RML structure to one reported for wildtype RML prion fibrils in an accompanying paper indicates that these post-translational modifications do not substantially alter the amyloid core conformation. This work demonstrates both common and divergent structural features of prion strains at the near-atomic level. |
 External links External links |  Nat Commun / Nat Commun /  PubMed:35831291 / PubMed:35831291 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 3.0 Å |
| Structure data | EMDB-25824, PDB-7td6: |
| Source |
|
 Keywords Keywords | PROTEIN FIBRIL / infectious prion / amyloid / parallel in-register |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers