+Search query
-Structure paper
| Title | The Molecular Basis for Apolipoprotein E4 as the Major Risk Factor for Late-Onset Alzheimer's Disease. |
|---|---|
| Journal, issue, pages | J Mol Biol, Vol. 431, Issue 12, Page 2248-2265, Year 2019 |
| Publish date | May 31, 2019 |
 Authors Authors | Ana-Caroline Raulin / Lucas Kraft / Youssra K Al-Hilaly / Wei-Feng Xue / John E McGeehan / John R Atack / Louise Serpell /  |
| PubMed Abstract | Apolipoprotein E4 (ApoE4) is one of three (E2, E3 and E4) human isoforms of an α-helical, 299-amino-acid protein. Homozygosity for the ε4 allele is the major genetic risk factor for developing late- ...Apolipoprotein E4 (ApoE4) is one of three (E2, E3 and E4) human isoforms of an α-helical, 299-amino-acid protein. Homozygosity for the ε4 allele is the major genetic risk factor for developing late-onset Alzheimer's disease (AD). ApoE2, ApoE3 and ApoE4 differ at amino acid positions 112 and 158, and these sequence variations may confer conformational differences that underlie their participation in the risk of developing AD. Here, we compared the shape, oligomerization state, conformation and stability of ApoE isoforms using a range of complementary biophysical methods including small-angle x-ray scattering, analytical ultracentrifugation, circular dichroism, x-ray fiber diffraction and transmission electron microscopy We provide an in-depth and definitive study demonstrating that all three proteins are similar in stability and conformation. However, we show that ApoE4 has a propensity to polymerize to form wavy filaments, which do not share the characteristics of cross-β amyloid fibrils. Moreover, we provide evidence for the inhibition of ApoE4 fibril formation by ApoE3. This study shows that recombinant ApoE isoforms show no significant differences at the structural or conformational level. However, self-assembly of the ApoE4 isoform may play a role in pathogenesis, and these results open opportunities for uncovering new triggers for AD onset. |
 External links External links |  J Mol Biol / J Mol Biol /  PubMed:31051176 / PubMed:31051176 /  PubMed Central PubMed Central |
| Methods | SAS (X-ray synchrotron) |
| Structure data | 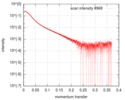 SASDD98: Apolipoprotein E3 (ApoE3) 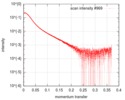 SASDDA8: Apolipoprotein E4 (ApoE4) 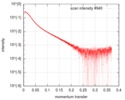 SASDDD7: Apolipoprotein E2 (ApoE2) |
| Source |
|
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers



 Homo sapiens (human)
Homo sapiens (human)