[English] 日本語
 Yorodumi
Yorodumi- EMDB-2033: Unsymmetrized reconstruction of the FLNa16-21 rod2 segment bound ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2033 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Unsymmetrized reconstruction of the FLNa16-21 rod2 segment bound with integrin Beta7 subunit tail | |||||||||
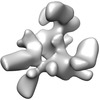
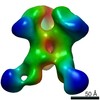
 Map data Map data | This is an unsymmetrized reconstruction of the filamin rod2 segment comprising domains 16-21 in the presence of a tenfold stoichiometric excess of integrin Beta7 subunit tail | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | cell adhesion / integrin signaling / signal transduction / single particle | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 15.7 Å | |||||||||
 Authors Authors | Hofmann GW / Jiang P / Campbell ID / Gilbert RJC | |||||||||
 Citation Citation |  Journal: Biochem J / Year: 2012 Journal: Biochem J / Year: 2012Title: The C-terminal rod 2 fragment of filamin A forms a compact structure that can be extended. Authors: Salla Ruskamo / Robert Gilbert / Gregor Hofmann / Pengju Jiang / Iain D Campbell / Jari Ylänne / Ulla Pentikäinen /  Abstract: Filamins are large proteins that cross-link actin filaments and connect to other cellular components. The C-terminal rod 2 region of FLNa (filamin A) mediates dimerization and interacts with several ...Filamins are large proteins that cross-link actin filaments and connect to other cellular components. The C-terminal rod 2 region of FLNa (filamin A) mediates dimerization and interacts with several transmembrane receptors and intracellular signalling adaptors. SAXS (small-angle X-ray scattering) experiments were used to make a model of a six immunoglobulin-like domain fragment of the FLNa rod 2 (domains 16-21). This fragment had a surprising three-branched structural arrangement, where each branch was made of a tightly packed two-domain pair. Peptides derived from transmembrane receptors and intracellular signalling proteins induced a more open structure of the six domain fragment. Mutagenesis studies suggested that these changes are caused by peptides binding to the CD faces on domains 19 and 21 which displace the preceding domain A-strands (18 and 20 respectively), thus opening the individual domain pairs. A single particle cryo-EM map of a nine domain rod 2 fragment (domains 16-24), showed a relatively compact dimeric particle and confirmed the three-branched arrangement as well as the peptide-induced conformation changes. These findings reveal features of filamin structure that are important for its interactions and mechanical properties. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2033.map.gz emd_2033.map.gz | 678 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2033-v30.xml emd-2033-v30.xml emd-2033.xml emd-2033.xml | 10.1 KB 10.1 KB | Display Display |  EMDB header EMDB header |
| Images |  EMD-2033.png EMD-2033.png | 65.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2033 http://ftp.pdbj.org/pub/emdb/structures/EMD-2033 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2033 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2033 | HTTPS FTP |
-Validation report
| Summary document |  emd_2033_validation.pdf.gz emd_2033_validation.pdf.gz | 196 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2033_full_validation.pdf.gz emd_2033_full_validation.pdf.gz | 195.1 KB | Display | |
| Data in XML |  emd_2033_validation.xml.gz emd_2033_validation.xml.gz | 5.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2033 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2033 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2033 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2033 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2033.map.gz / Format: CCP4 / Size: 3.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2033.map.gz / Format: CCP4 / Size: 3.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is an unsymmetrized reconstruction of the filamin rod2 segment comprising domains 16-21 in the presence of a tenfold stoichiometric excess of integrin Beta7 subunit tail | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.37 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Filamin A rod-2 segment, comprising domains 16-21 bound with an i...
| Entire | Name: Filamin A rod-2 segment, comprising domains 16-21 bound with an integrin Beta7 subunit tail |
|---|---|
| Components |
|
-Supramolecule #1000: Filamin A rod-2 segment, comprising domains 16-21 bound with an i...
| Supramolecule | Name: Filamin A rod-2 segment, comprising domains 16-21 bound with an integrin Beta7 subunit tail type: sample / ID: 1000 / Oligomeric state: Dimer / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 200 KDa / Theoretical: 200 KDa |
-Macromolecule #1: FLNa16-21-Beta7 tail
| Macromolecule | Name: FLNa16-21-Beta7 tail / type: protein_or_peptide / ID: 1 / Name.synonym: Activated filamin A rod 2 Details: The sample was imaged as a monodisperse distribution in vitreous ice Number of copies: 2 / Oligomeric state: Dimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human / Location in cell: Cytoplasm (adjacent plasma membrane) Homo sapiens (human) / synonym: Human / Location in cell: Cytoplasm (adjacent plasma membrane) |
| Molecular weight | Experimental: 200 KDa / Theoretical: 200 KDa |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL |
|---|---|
| Grid | Details: 300 mesh copper with lacey carbon |
| Vitrification | Cryogen name: ETHANE / Instrument: HOMEMADE PLUNGER / Details: Vitrification instrument: Home made guillotine Method: Whatman No. 1 paper blot for 1-2 seconds prior to plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Temperature | Average: 110 K |
| Alignment procedure | Legacy - Astigmatism: At 112,000 times magnification |
| Date | Sep 1, 2006 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 7 µm / Number real images: 48 / Od range: 5 / Bits/pixel: 8 |
| Tilt angle min | 0 |
| Tilt angle max | 0 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 59000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | The raw images were treated with a three-level decomposition with a fourth-order bi-orthogonal wavelet function in MATLAB to enhance contrast. Particles were selected semi-automatically using Boxer. |
|---|---|
| CTF correction | Details: per micrograph |
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 15.7 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: IMAGIC, SPIDER Details: The fineness of sampling was restricted to 15 degrees to reflect inaccuracy of angle determination at this particle size at finer spacings. The resolution of the map was therefore fixed at 30 Angstrom. Number images used: 32477 |
| Final angle assignment | Details: SPIDER Euler angles. |
 Movie
Movie Controller
Controller