[English] 日本語
 Yorodumi
Yorodumi- EMDB-9839: Structure of PSII-FCP supercomplex from a centric diatom Chaetoce... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9839 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of PSII-FCP supercomplex from a centric diatom Chaetoceros gracilis at 3.02 angstrom resolution | |||||||||
 Map data Map data | PSII-FCP | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | PSII-FCP supercomplex / PLANT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationphotosystem II assembly / photosystem II stabilization / photosystem II / extrinsic component of membrane / phosphate ion binding / photosynthesis, light reaction / chloroplast thylakoid membrane / photosynthesis / respiratory electron transport chain / electron transfer activity ...photosystem II assembly / photosystem II stabilization / photosystem II / extrinsic component of membrane / phosphate ion binding / photosynthesis, light reaction / chloroplast thylakoid membrane / photosynthesis / respiratory electron transport chain / electron transfer activity / protein stabilization / iron ion binding / heme binding Similarity search - Function | |||||||||
| Biological species |  Chaetoceros gracilis (Diatom) Chaetoceros gracilis (Diatom) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.02 Å | |||||||||
 Authors Authors | Pi X / Zhao S | |||||||||
 Citation Citation |  Journal: Science / Year: 2019 Journal: Science / Year: 2019Title: The pigment-protein network of a diatom photosystem II-light-harvesting antenna supercomplex. Authors: Xiong Pi / Songhao Zhao / Wenda Wang / Desheng Liu / Caizhe Xu / Guangye Han / Tingyun Kuang / Sen-Fang Sui / Jian-Ren Shen /   Abstract: Diatoms play important roles in global primary productivity and biogeochemical cycling of carbon, in part owing to the ability of their photosynthetic apparatus to adapt to rapidly changing light ...Diatoms play important roles in global primary productivity and biogeochemical cycling of carbon, in part owing to the ability of their photosynthetic apparatus to adapt to rapidly changing light intensity. We report a cryo-electron microscopy structure of the photosystem II (PSII)-fucoxanthin (Fx) chlorophyll (Chl) a/c binding protein (FCPII) supercomplex from the centric diatom The supercomplex comprises two protomers, each with two tetrameric and three monomeric FCPIIs around a PSII core that contains five extrinsic oxygen-evolving proteins at the lumenal surface. The structure reveals the arrangement of a huge pigment network that contributes to efficient light energy harvesting, transfer, and dissipation processes in the diatoms. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9839.map.gz emd_9839.map.gz | 138.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9839-v30.xml emd-9839-v30.xml emd-9839.xml emd-9839.xml | 52.2 KB 52.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9839.png emd_9839.png | 188.3 KB | ||
| Filedesc metadata |  emd-9839.cif.gz emd-9839.cif.gz | 11.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9839 http://ftp.pdbj.org/pub/emdb/structures/EMD-9839 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9839 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9839 | HTTPS FTP |
-Related structure data
| Related structure data |  6jluMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9839.map.gz / Format: CCP4 / Size: 149.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9839.map.gz / Format: CCP4 / Size: 149.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | PSII-FCP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.30654 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
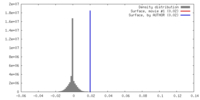
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : PSII-FCP supercomplex
+Supramolecule #1: PSII-FCP supercomplex
+Macromolecule #1: PsbA
+Macromolecule #2: PsbB
+Macromolecule #3: PsbC
+Macromolecule #4: PsbD
+Macromolecule #5: PsbE
+Macromolecule #6: PsbF
+Macromolecule #7: Psb31
+Macromolecule #8: Photosystem II reaction center protein H
+Macromolecule #9: PsbI
+Macromolecule #10: PsbJ
+Macromolecule #11: PsbK
+Macromolecule #12: PsbL
+Macromolecule #13: PsbM
+Macromolecule #14: Psb34
+Macromolecule #15: Extrinsic protein in photosystem II
+Macromolecule #16: FCP-D
+Macromolecule #17: Extrinsic protein in photosystem II
+Macromolecule #18: PsbG
+Macromolecule #19: PsbT
+Macromolecule #20: Extrinsic protein in photosystem II
+Macromolecule #21: Cytochrome c-550
+Macromolecule #22: PsbW
+Macromolecule #23: Photosystem II reaction center X protein
+Macromolecule #24: PsbY
+Macromolecule #25: PsbZ
+Macromolecule #26: FCP-E
+Macromolecule #27: FCP-A
+Macromolecule #28: FCP-F
+Macromolecule #29: CA-MN4-O5 CLUSTER
+Macromolecule #30: FE (II) ION
+Macromolecule #31: CHLOROPHYLL A
+Macromolecule #32: PHEOPHYTIN A
+Macromolecule #33: BETA-CAROTENE
+Macromolecule #34: 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL
+Macromolecule #35: 2,3-DIMETHYL-5-(3,7,11,15,19,23,27,31,35-NONAMETHYL-2,6,10,14,18,...
+Macromolecule #36: 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE
+Macromolecule #37: 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE
+Macromolecule #38: DIGALACTOSYL DIACYL GLYCEROL (DGDG)
+Macromolecule #39: CHLORIDE ION
+Macromolecule #40: BICARBONATE ION
+Macromolecule #41: PROTOPORPHYRIN IX CONTAINING FE
+Macromolecule #42: Chlorophyll c1
+Macromolecule #43: (3S,3'S,5R,5'R,6S,6'R,8'R)-3,5'-dihydroxy-8-oxo-6',7'-didehydro-5...
+Macromolecule #44: (3S,3'R,5R,6S,7cis)-7',8'-didehydro-5,6-dihydro-5,6-epoxy-beta,be...
+Macromolecule #45: Chlorophyll c2
+Macromolecule #46: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 6.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)