[English] 日本語
 Yorodumi
Yorodumi- EMDB-8515: Chicken Slo2.2 in an open conformation vitrified in the presence ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8515 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Chicken Slo2.2 in an open conformation vitrified in the presence of 300 mM NaCl | |||||||||
 Map data Map data | Open Slo2.2 density map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ion channel / potassium channel / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationintracellular sodium-activated potassium channel activity / outward rectifier potassium channel activity / potassium ion transmembrane transport / protein homotetramerization / metal ion binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.76 Å | |||||||||
 Authors Authors | Hite RK / MacKinnon R | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2017 Journal: Cell / Year: 2017Title: Structural Titration of Slo2.2, a Na-Dependent K Channel. Authors: Richard K Hite / Roderick MacKinnon /  Abstract: The stable structural conformations that occur along the complete reaction coordinate for ion channel opening have never been observed. In this study, we describe the equilibrium ensemble of ...The stable structural conformations that occur along the complete reaction coordinate for ion channel opening have never been observed. In this study, we describe the equilibrium ensemble of structures of Slo2.2, a neuronal Na-activated K channel, as a function of the Na concentration. We find that Slo2.2 exists in multiple closed conformations whose relative occupancies are independent of Na concentration. An open conformation emerges from an ensemble of closed conformations in a highly Na-dependent manner, without evidence of Na-dependent intermediates. In other words, channel opening is a highly concerted, switch-like process. The midpoint of the structural titration matches that of the functional titration. A maximum open conformation probability approaching 1.0 and maximum functional open probability approaching 0.7 imply that, within the class of open channels, there is a subclass that is not permeable to ions. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8515.map.gz emd_8515.map.gz | 37.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8515-v30.xml emd-8515-v30.xml emd-8515.xml emd-8515.xml | 15.4 KB 15.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_8515.png emd_8515.png | 172.8 KB | ||
| Filedesc metadata |  emd-8515.cif.gz emd-8515.cif.gz | 6.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8515 http://ftp.pdbj.org/pub/emdb/structures/EMD-8515 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8515 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8515 | HTTPS FTP |
-Related structure data
| Related structure data |  5u70MC  8517C  5u76C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8515.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8515.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Open Slo2.2 density map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.3 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
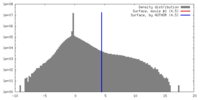
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Chicken Slo2.2 in open conformation
| Entire | Name: Chicken Slo2.2 in open conformation |
|---|---|
| Components |
|
-Supramolecule #1: Chicken Slo2.2 in open conformation
| Supramolecule | Name: Chicken Slo2.2 in open conformation / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Potassium channel subfamily T member 1
| Macromolecule | Name: Potassium channel subfamily T member 1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 137.409812 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MARAKLKNSP SESNSHVKTV PPATTEDVRG VSPLLPARRM GSLGSDVGQR PHAEDFSMDS SFSQVQVEFY VNENTFKERL KLFFIKNQR SSLRIRLFNF SLKLLTCLLY IVRVLLDNPE EGIGCWECEK QNYTLFNQST KINWSHIFWV DRKLPLWAVQ V SIALISFL ...String: MARAKLKNSP SESNSHVKTV PPATTEDVRG VSPLLPARRM GSLGSDVGQR PHAEDFSMDS SFSQVQVEFY VNENTFKERL KLFFIKNQR SSLRIRLFNF SLKLLTCLLY IVRVLLDNPE EGIGCWECEK QNYTLFNQST KINWSHIFWV DRKLPLWAVQ V SIALISFL ETMLLIYLSY KGNIWEQIFR ISFILEMINT VPFIITIFWP PLRNLFIPVF LNCWLAKYAL ENMINDLHRA IQ RTQSAMF NQVLILICTL LCLVFTGTCG IQHLERAGEK LSLFKSFYFC IVTFSTVGYG DVTPKIWPSQ LLVVIMICVA LVV LPLQFE ELVYLWMERQ KSGGNYSRHR AQTEKHVVLC VSSLKIDLLM DFLNEFYAHP RLQDYYVVIL CPTEMDIQVR RVLQ IPLWS QRVIYLQGSA LKDQDLMRAK MDNGEACFIL SSRNEVDRTA ADHQTILRAW AVKDFAPNCP LYVQILKPEN KFHVK FADH VVCEEECKYA MLALNCVCPA TSTLITLLVH TSRGQEGQES PEQWQRMYGR CSGNEVYHIR MGDSKFFMEY EGKSFT YAA FHAHKKYGVC LIGIRREENK SILLNPGPRH IMAASDTCFY INITKEENSA FIFKQAEKQK KKGFAGRGTY DGPSRLP VH SIIASMGTVA MDLQNTECRP TNSSKLALPA ENGSGNRRPS IAPVLELADT SSLLPCDLLS DQSEDEMTQS DEEGSAVV E YVKGYPPNSP YIGSSPTLCH LLPEKAPFCC LRLDKGCKHN SFEDAKAYGF KNKLIIVSAE TAGNGLYNFI VPLRAYYRS RKELNPIVLL LDNKPEHHFL EAICCFPMVY YMEGTIDNLD SLLQCGIIYA DNLVVVDKES TMSAEEDYMA DAKTIVNVQT MFRLFPSLS IITELTHPSN MRFMQFRAKD SYSLALSKLE KKERENGSNL AFMFRLPFAA GRVFSISMLD TLLYQSFVKD Y MITITRLL LGLDTTPGSG YLCAMKITED DLWIRTYGRL FQKLCSSSAE IPIGIYRTES HMFATSEPHD IRAQSQISIN VE DCEDTKD VKEHWGIKTG HHRNSCSSDQ SEHPLLRRKS MQWARRLSRK GNKHSGKTAE WISQQRLSLY RRSERQELSE LVK NRMKHL GLPTTGYDEM NDHQNTLSYV LINPPPDTRL ELNDIVYLIR SDPLAHVAND GHSRKSSCSN KLGPCNPETR DETQ L UniProtKB: Potassium channel subfamily T member 1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 6 mg/mL | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 10 sec. | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 88 % / Chamber temperature: 298 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Dimensions - Width: 3710 pixel / Digitization - Dimensions - Height: 3838 pixel / Digitization - Frames/image: 1-50 / Number grids imaged: 1 / Number real images: 2000 / Average exposure time: 1.2 sec. / Average electron dose: 59.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-5u70: |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)