[English] 日本語
 Yorodumi
Yorodumi- EMDB-7000: Subtomogram average of microtubule-bound dynein-dynactin-BICD2N c... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7000 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Subtomogram average of microtubule-bound dynein-dynactin-BICD2N complex | |||||||||
 Map data Map data | Microtubule-bound dynein-dynactin-BICD2 complex | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 38.0 Å | |||||||||
 Authors Authors | Grotjahn DA / Chowdhury S / Xu Y / McKenney RJ / Schroer TA / Lander GC | |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2018 Journal: Nat Struct Mol Biol / Year: 2018Title: Cryo-electron tomography reveals that dynactin recruits a team of dyneins for processive motility. Authors: Danielle A Grotjahn / Saikat Chowdhury / Yiru Xu / Richard J McKenney / Trina A Schroer / Gabriel C Lander /  Abstract: Cytoplasmic dynein is a protein complex that transports molecular cargo along microtubules (MTs), playing a key role in the intracellular trafficking network. Vertebrate dynein's movement becomes ...Cytoplasmic dynein is a protein complex that transports molecular cargo along microtubules (MTs), playing a key role in the intracellular trafficking network. Vertebrate dynein's movement becomes strikingly enhanced upon interacting with dynactin and a cargo adaptor such as BicaudalD2. However, the mechanisms responsible for increased transport activity are not well understood, largely owing to limited structural information. We used cryo-electron tomography (cryo-ET) to visualize the 3D structure of the MT-bound dynein-dynactin complex from Mus musculus and show that the dynactin-cargo adaptor complex binds two dimeric dyneins. This configuration imposes spatial and conformational constraints on both dynein dimers, positioning the four motor domains in proximity to one another and oriented toward the MT minus end. We propose that grouping multiple dyneins onto a single dynactin scaffold promotes collective force production, increased processivity, and unidirectional movement, suggesting mechanistic parallels to axonemal dynein. These findings provide structural insights into a previously unknown mechanism for dynein regulation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7000.map.gz emd_7000.map.gz | 2.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7000-v30.xml emd-7000-v30.xml emd-7000.xml emd-7000.xml | 19.3 KB 19.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7000.png emd_7000.png | 94.7 KB | ||
| Others |  emd_7000_additional_1.map.gz emd_7000_additional_1.map.gz emd_7000_additional_2.map.gz emd_7000_additional_2.map.gz emd_7000_additional_3.map.gz emd_7000_additional_3.map.gz | 2.4 MB 2.3 MB 2.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7000 http://ftp.pdbj.org/pub/emdb/structures/EMD-7000 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7000 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7000 | HTTPS FTP |
-Validation report
| Summary document |  emd_7000_validation.pdf.gz emd_7000_validation.pdf.gz | 79.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_7000_full_validation.pdf.gz emd_7000_full_validation.pdf.gz | 78.3 KB | Display | |
| Data in XML |  emd_7000_validation.xml.gz emd_7000_validation.xml.gz | 493 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7000 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7000 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7000 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-7000 | HTTPS FTP |
-Related structure data
| Related structure data |  7001C C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10520 (Title: Cryo-electron tomography reveals that dynactin recruits a team of dyneins for processive motility. EMPIAR-10520 (Title: Cryo-electron tomography reveals that dynactin recruits a team of dyneins for processive motility.Data size: 200.1 / Data #1: tilt series in mrc format [tilt series]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_7000.map.gz / Format: CCP4 / Size: 3.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7000.map.gz / Format: CCP4 / Size: 3.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Microtubule-bound dynein-dynactin-BICD2 complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 8.52 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
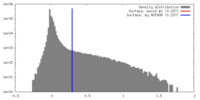
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: focused reconstruction of dynein dimer-2 motor domains
| File | emd_7000_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | focused reconstruction of dynein dimer-2 motor domains | ||||||||||||
| Projections & Slices |
| ||||||||||||
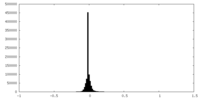
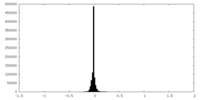
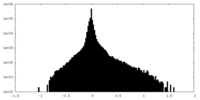
| Density Histograms |
-Additional map: focused reconstruction of dynein tails associated with dynactin-BICD2N...
| File | emd_7000_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | focused reconstruction of dynein tails associated with dynactin-BICD2N | ||||||||||||
| Projections & Slices |
| ||||||||||||
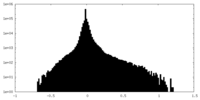
| Density Histograms |
-Additional map: focused reconstruction of dynein dimer-1 motor domains
| File | emd_7000_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | focused reconstruction of dynein dimer-1 motor domains | ||||||||||||
| Projections & Slices |
| ||||||||||||
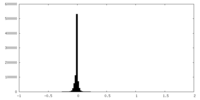
| Density Histograms |
- Sample components
Sample components
-Entire : microtubule-bound dynein-dynactin-BICD2N complex
| Entire | Name: microtubule-bound dynein-dynactin-BICD2N complex |
|---|---|
| Components |
|
-Supramolecule #1: microtubule-bound dynein-dynactin-BICD2N complex
| Supramolecule | Name: microtubule-bound dynein-dynactin-BICD2N complex / type: complex / ID: 1 / Parent: 0 Details: GFP-BICD2N fragment recombinantly expressed and purified. Microtubule-bound dynein-dynactin complexes isolated from mouse brain tissue. |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 4.2 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 Details: PMEE buffer (35 mM PIPES, 5 mM MgSO4, 1 mM EGTA, 0.5 mM EDTA, 1 mM GTP, 4 mM AMPPNP, 20 uM Taxol) |
|---|---|
| Grid | Model: Quantifoil, UltrAuFoil, R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Atmosphere: OTHER |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 85 % / Chamber temperature: 277.15 K / Instrument: HOMEMADE PLUNGER Details: 4 uL of sample was applied to one side of a plasma-cleaned grid, and the grid was manually blotted on the opposite side for 5-7 seconds immediately before vitrification.. |
| Details | Microtubule-bound dynein-dynactin-BICD2N complexes isolated from mouse brain tissue in the presence of non-hydrolyzable ATP analog AMPPNP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 98.15 K / Max: 98.15 K |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3710 pixel / Digitization - Dimensions - Height: 3838 pixel / Digitization - Sampling interval: 5.0 µm / Digitization - Frames/image: 1-15 / Number grids imaged: 1 / Average exposure time: 1.16 sec. / Average electron dose: 0.95 e/Å2 Details: Images were collected in movie mode with a per-frame exposure time of 80 ms. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated magnification: 14000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 8.0 µm / Nominal defocus min: 6.0 µm / Nominal magnification: 14000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | Movie frames per tilt were aligned using MotionCorr. |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: FOURIER SPACE / Resolution.type: BY AUTHOR / Resolution: 38.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: RELION (ver. 1.4) Details: Initial angular assignments were obtained by manually docking available structures/reconstructions into extracted subvolumes using UCSF Chimera. These were further refined using the RELION 1. ...Details: Initial angular assignments were obtained by manually docking available structures/reconstructions into extracted subvolumes using UCSF Chimera. These were further refined using the RELION 1.4 subtomogram averaging auto-refinement program, followed by focused refinement of individual subregions of the complex by application of binary masks and continuation of refinement. The resulting focused maps were stitched together using the "vop maximum" function in UCSF Chimera. Number subtomograms used: 502 |
| Extraction | Number tomograms: 127 / Number images used: 502 / Method: manual / Software - Name: EMAN2 (ver. 2.11) / Software - details: e2spt_boxer Details: Subvolumes were picked using the EMAN2 manual picker for subvolumes. |
| Final angle assignment | Type: NOT APPLICABLE / Software - Name: RELION (ver. 1.4) / Software - details: Subtomogram Averaging |
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT |
|---|
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)