+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of kisspeptin receptor bound to TAK-448 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cryo-EM / kisspeptin receptor / GPR54 / G proteins / MEMBRANE PROTEIN / KISS1R | |||||||||
| Function / homology |  Function and homology information Function and homology informationFatty Acids bound to GPR40 (FFAR1) regulate insulin secretion / Acetylcholine regulates insulin secretion / neuropeptide receptor activity / sensory perception of itch / phospholipase C-activating G protein-coupled glutamate receptor signaling pathway / PLC beta mediated events / phospholipase C-activating serotonin receptor signaling pathway / regulation of platelet activation / entrainment of circadian clock / positive regulation of hormone secretion ...Fatty Acids bound to GPR40 (FFAR1) regulate insulin secretion / Acetylcholine regulates insulin secretion / neuropeptide receptor activity / sensory perception of itch / phospholipase C-activating G protein-coupled glutamate receptor signaling pathway / PLC beta mediated events / phospholipase C-activating serotonin receptor signaling pathway / regulation of platelet activation / entrainment of circadian clock / positive regulation of hormone secretion / regulation of canonical Wnt signaling pathway / glutamate receptor signaling pathway / mast cell degranulation / phototransduction, visible light / photoreceptor outer segment / postsynaptic cytosol / cellular response to acidic pH / adenylate cyclase inhibitor activity / positive regulation of protein localization to cell cortex / T cell migration / positive regulation of relaxation of smooth muscle / Adenylate cyclase inhibitory pathway / hormone-mediated signaling pathway / D2 dopamine receptor binding / adenylate cyclase-inhibiting serotonin receptor signaling pathway / G protein-coupled serotonin receptor binding / cellular response to forskolin / Turbulent (oscillatory, disturbed) flow shear stress activates signaling by PIEZO1 and integrins in endothelial cells / Peptide ligand-binding receptors / regulation of mitotic spindle organization / chemokine-mediated signaling pathway / GTPase activator activity / Regulation of insulin secretion / neuropeptide signaling pathway / response to prostaglandin E / positive regulation of cholesterol biosynthetic process / negative regulation of insulin secretion / G protein-coupled receptor binding / response to peptide hormone / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / G-protein beta/gamma-subunit complex binding / centriolar satellite / blood coagulation / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / Glucagon signaling in metabolic regulation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / photoreceptor disc membrane / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / GDP binding / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / cellular response to catecholamine stimulus / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / cellular response to prostaglandin E stimulus / heterotrimeric G-protein complex / G alpha (12/13) signalling events / Inactivation, recovery and regulation of the phototransduction cascade / G-protein beta-subunit binding / extracellular vesicle / sensory perception of taste / adenylate cyclase-activating G protein-coupled receptor signaling pathway / sperm principal piece / Thrombin signalling through proteinase activated receptors (PARs) / signaling receptor complex adaptor activity / retina development in camera-type eye / GTPase binding / G protein activity / fibroblast proliferation / Ca2+ pathway / midbody / cell cortex / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / nuclear membrane / G alpha (i) signalling events / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (q) signalling events / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / Ras protein signal transduction Similarity search - Function | |||||||||
| Biological species |  Homo Sapiens (human) / Homo Sapiens (human) /  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.07 Å | |||||||||
 Authors Authors | Shen S / Liu H / Xu HE | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2024 Journal: Cell Rep / Year: 2024Title: Structural basis for hormone recognition and distinctive Gq protein coupling by the kisspeptin receptor. Authors: Shiyi Shen / Dongxue Wang / Heng Liu / Xinheng He / Yinglong Cao / Juanhua Chen / Shujie Li / Xi Cheng / H Eric Xu / Jia Duan /  Abstract: Kisspeptin signaling through its G protein-coupled receptor, KISS1R, plays an indispensable role in regulating reproduction via the hypothalamic-pituitary-gonadal axis. Dysregulation of this pathway ...Kisspeptin signaling through its G protein-coupled receptor, KISS1R, plays an indispensable role in regulating reproduction via the hypothalamic-pituitary-gonadal axis. Dysregulation of this pathway underlies severe disorders like infertility and precocious puberty. Here, we present cryo-EM structures of KISS1R bound to the endogenous agonist kisspeptin-10 and a synthetic analog TAK-448. These structures reveal pivotal interactions between peptide ligands and KISS1R extracellular loops for receptor activation. Both peptides exhibit a conserved binding mode, unveiling their common activation mechanism. Intriguingly, KISS1R displays a distinct 40° angular deviation in its intracellular TM6 region compared to other G-coupled receptors, enabling distinct interactions with G. This study reveals the molecular intricacies governing ligand binding and activation of KISS1R, while highlighting its exceptional ability to couple with G. Our findings pave the way for structure-guided design of therapeutics targeting this physiologically indispensable receptor. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_60142.map.gz emd_60142.map.gz | 59.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-60142-v30.xml emd-60142-v30.xml emd-60142.xml emd-60142.xml | 19.5 KB 19.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_60142.png emd_60142.png | 51.1 KB | ||
| Masks |  emd_60142_msk_1.map emd_60142_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-60142.cif.gz emd-60142.cif.gz | 6.9 KB | ||
| Others |  emd_60142_half_map_1.map.gz emd_60142_half_map_1.map.gz emd_60142_half_map_2.map.gz emd_60142_half_map_2.map.gz | 59.3 MB 59.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-60142 http://ftp.pdbj.org/pub/emdb/structures/EMD-60142 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60142 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60142 | HTTPS FTP |
-Related structure data
| Related structure data |  8zjeMC  8zjdC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_60142.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_60142.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.73 Å | ||||||||||||||||||||||||||||||||||||
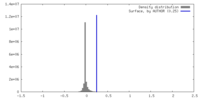
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_60142_msk_1.map emd_60142_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_60142_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_60142_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : cryo-EM structure of kisspeptin receptor in complex with G proteins
| Entire | Name: cryo-EM structure of kisspeptin receptor in complex with G proteins |
|---|---|
| Components |
|
-Supramolecule #1: cryo-EM structure of kisspeptin receptor in complex with G proteins
| Supramolecule | Name: cryo-EM structure of kisspeptin receptor in complex with G proteins type: complex / ID: 1 / Parent: 0 / Macromolecule list: #2-#5, #1 |
|---|---|
| Source (natural) | Organism:  Homo Sapiens (human) Homo Sapiens (human) |
-Macromolecule #1: Guanine nucleotide-binding protein G(i) subunit alpha-1,Guanine n...
| Macromolecule | Name: Guanine nucleotide-binding protein G(i) subunit alpha-1,Guanine nucleotide-binding protein G(q) subunit alpha type: protein_or_peptide / ID: 1 Details: fusion protein of Guanine nucleotide-binding protein G(i) subunit alpha-1 and Guanine nucleotide-binding protein G(q) subunit alpha-q. Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 41.312922 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGCTLSAEDK AAVERSKMID RNLREDGERS RRELKLLLLG TGESGKSTFI KQMRIIHGSG YSDEDKRGFT KLVYQNIFTA MQAMIRAMD TLKIPYKYEH NKAHAQLVRE VDVEKVSAFE NPYVDAIKSL WNDPGIQECY DRRREYQLSD STKYYLNDLD R VADPAYLP ...String: MGCTLSAEDK AAVERSKMID RNLREDGERS RRELKLLLLG TGESGKSTFI KQMRIIHGSG YSDEDKRGFT KLVYQNIFTA MQAMIRAMD TLKIPYKYEH NKAHAQLVRE VDVEKVSAFE NPYVDAIKSL WNDPGIQECY DRRREYQLSD STKYYLNDLD R VADPAYLP TQQDVLRVRV PTTGIIEYPF DLQSVIFRMV DVGAQRSERR KWIHCFENVT SIMFLVALSE YDQVLVESDN EN RMEESKA LFRTIITYPW FQNSSVILFL NKKDLLEEKI MYSHLVDYFP EYDGPQRDAQ AAREFILKMF VDLNPDSDKI IYS HFTCST DTENIRFVFA AVKDTILQLN LKEYNLV UniProtKB: Guanine nucleotide-binding protein G(i) subunit alpha-1, Guanine nucleotide-binding protein G(q) subunit alpha |
-Macromolecule #2: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 38.744371 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MHHHHHHGSL LQSELDQLRQ EAEQLKNQIR DARKACADAT LSQITNNIDP VGRIQMRTRR TLRGHLAKIY AMHWGTDSRL LVSASQDGK LIIWDSYTTN KVHAIPLRSS WVMTCAYAPS GNYVACGGLD NICSIYNLKT REGNVRVSRE LAGHTGYLSC C RFLDDNQI ...String: MHHHHHHGSL LQSELDQLRQ EAEQLKNQIR DARKACADAT LSQITNNIDP VGRIQMRTRR TLRGHLAKIY AMHWGTDSRL LVSASQDGK LIIWDSYTTN KVHAIPLRSS WVMTCAYAPS GNYVACGGLD NICSIYNLKT REGNVRVSRE LAGHTGYLSC C RFLDDNQI VTSSGDTTCA LWDIETGQQT TTFTGHTGDV MSLSLAPDTR LFVSGACDAS AKLWDVREGM CRQTFTGHES DI NAICFFP NGNAFATGSD DATCRLFDLR ADQELMTYSH DNIICGITSV SFSKSGRLLL AGYDDFNCNV WDALKADRAG VLA GHDNRV SCLGVTDDGM AVATGSWDSF LKIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #4: KiSS-1 receptor
| Macromolecule | Name: KiSS-1 receptor / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 42.747094 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MHTVATSGPN ASWGAPANAS GCPGCGANAS DGPVPSPRAV DAWLVPLFFA ALMLLGLVGN SLVIYVICRH KPMRTVTNFY IANLAATDV TFLLCCVPFT ALLYPLPGWV LGDFMCKFVN YIQQVSVQAT CWTLTAMSVD RWYVTVFPLR ALHRRTPRLA L AVSLSIWV ...String: MHTVATSGPN ASWGAPANAS GCPGCGANAS DGPVPSPRAV DAWLVPLFFA ALMLLGLVGN SLVIYVICRH KPMRTVTNFY IANLAATDV TFLLCCVPFT ALLYPLPGWV LGDFMCKFVN YIQQVSVQAT CWTLTAMSVD RWYVTVFPLR ALHRRTPRLA L AVSLSIWV GSAAVSAPVL ALHRLSPGPR AYCSEAFPSR ALERAFALYN LLALYLLPLL ATCACYAAML RHLGRVAVRP AP ADSALQG QVLAERAGAV RAKVSRLVAA VVLLFAACWG PIQLFLVLQA LGPAGSWHPR SYAAYALKTW AHCMSYSNSA LNP LLYAFL GSHFRQAFRR VCPCAPRRPR RPRRPGPSDP AAPHAELLRL GSHPAPARAQ KPGSSGLAAR GLCVLGEDNA PL UniProtKB: KiSS-1 receptor |
-Macromolecule #5: scFv16
| Macromolecule | Name: scFv16 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 26.277299 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: VQLVESGGGL VQPGGSRKLS CSASGFAFSS FGMHWVRQAP EKGLEWVAYI SSGSGTIYYA DTVKGRFTIS RDDPKNTLFL QMTSLRSED TAMYYCVRSI YYYGSSPFDF WGQGTTLTVS AGGGGSGGGG SGGGGSADIV MTQATSSVPV TPGESVSISC R SSKSLLHS ...String: VQLVESGGGL VQPGGSRKLS CSASGFAFSS FGMHWVRQAP EKGLEWVAYI SSGSGTIYYA DTVKGRFTIS RDDPKNTLFL QMTSLRSED TAMYYCVRSI YYYGSSPFDF WGQGTTLTVS AGGGGSGGGG SGGGGSADIV MTQATSSVPV TPGESVSISC R SSKSLLHS NGNTYLYWFL QRPGQSPQLL IYRMSNLASG VPDRFSGSGS GTAFTLTISR LEAEDVGVYY CMQHLEYPLT FG AGTKLEL |
-Macromolecule #6: ACE-DTY-HYP-ASN-THR-PHE-XZA-LEU-NMM-TRP-NH2
| Macromolecule | Name: ACE-DTY-HYP-ASN-THR-PHE-XZA-LEU-NMM-TRP-NH2 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 1.20835 KDa |
| Sequence | String: (ACE)(DTY)(HYP)NTF(XZA)L(NMM)W (NH2) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: DARK FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.07 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 111547 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller



































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)