[English] 日本語
 Yorodumi
Yorodumi- EMDB-47120: E. coli RNA polymerase consensus volume with a bound fluoride rib... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | E. coli RNA polymerase consensus volume with a bound fluoride riboswitch in the ligand-bound state | |||||||||
 Map data Map data | E. coli RNA polymerase consensus volume with a bound fluoride riboswitch in the liganded-bound state. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RNA polymerase / riboswitch / fluoride / TRANSFERASE-DNA-RNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationsubmerged biofilm formation / cellular response to cell envelope stress / cytosolic DNA-directed RNA polymerase complex / regulation of DNA-templated transcription initiation / bacterial-type flagellum assembly / bacterial-type flagellum-dependent cell motility / nitrate assimilation / DNA-directed RNA polymerase complex / transcription elongation factor complex / regulation of DNA-templated transcription elongation ...submerged biofilm formation / cellular response to cell envelope stress / cytosolic DNA-directed RNA polymerase complex / regulation of DNA-templated transcription initiation / bacterial-type flagellum assembly / bacterial-type flagellum-dependent cell motility / nitrate assimilation / DNA-directed RNA polymerase complex / transcription elongation factor complex / regulation of DNA-templated transcription elongation / transcription antitermination / DNA-templated transcription initiation / cell motility / ribonucleoside binding / DNA-directed 5'-3' RNA polymerase activity / DNA-directed RNA polymerase / response to heat / intracellular iron ion homeostasis / protein dimerization activity / response to antibiotic / DNA-templated transcription / magnesium ion binding / DNA binding / zinc ion binding / membrane / cytosol / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
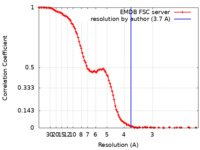
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | |||||||||
 Authors Authors | Porta JC / Ellinger E / Liu Y / Walter NG | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structural basis of RNA-mediated regulation of transcriptional pausing. Authors: Porta JC / Ellinger E / Liu Y / Chauvier A / Walter NG | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_47120.map.gz emd_47120.map.gz | 97.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-47120-v30.xml emd-47120-v30.xml emd-47120.xml emd-47120.xml | 26.3 KB 26.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_47120_fsc.xml emd_47120_fsc.xml | 13.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_47120.png emd_47120.png | 85.1 KB | ||
| Filedesc metadata |  emd-47120.cif.gz emd-47120.cif.gz | 8.7 KB | ||
| Others |  emd_47120_half_map_1.map.gz emd_47120_half_map_1.map.gz emd_47120_half_map_2.map.gz emd_47120_half_map_2.map.gz | 95.5 MB 95.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-47120 http://ftp.pdbj.org/pub/emdb/structures/EMD-47120 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-47120 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-47120 | HTTPS FTP |
-Validation report
| Summary document |  emd_47120_validation.pdf.gz emd_47120_validation.pdf.gz | 952.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_47120_full_validation.pdf.gz emd_47120_full_validation.pdf.gz | 951.9 KB | Display | |
| Data in XML |  emd_47120_validation.xml.gz emd_47120_validation.xml.gz | 16.8 KB | Display | |
| Data in CIF |  emd_47120_validation.cif.gz emd_47120_validation.cif.gz | 22.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-47120 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-47120 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-47120 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-47120 | HTTPS FTP |
-Related structure data
| Related structure data |  9dr1MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_47120.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_47120.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | E. coli RNA polymerase consensus volume with a bound fluoride riboswitch in the liganded-bound state. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1 Å | ||||||||||||||||||||||||||||||||||||
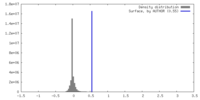
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half-map B of the E. coli RNA polymerase...
| File | emd_47120_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map B of the E. coli RNA polymerase consensus volume with a bound fluoride riboswitch in the ligand-bound state. | ||||||||||||
| Projections & Slices |
| ||||||||||||
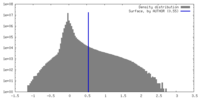
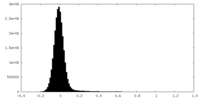
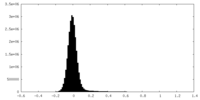
| Density Histograms |
-Half map: Half-map A of the E. coli RNA polymerase...
| File | emd_47120_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map A of the E. coli RNA polymerase consensus volume with a bound fluoride riboswitch in the ligand-bound state. | ||||||||||||
| Projections & Slices |
| ||||||||||||
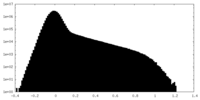
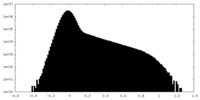
| Density Histograms |
- Sample components
Sample components
-Entire : E. coli RNA polymerase with a bound fluoride riboswitch in the un...
| Entire | Name: E. coli RNA polymerase with a bound fluoride riboswitch in the unliganded state |
|---|---|
| Components |
|
-Supramolecule #1: E. coli RNA polymerase with a bound fluoride riboswitch in the un...
| Supramolecule | Name: E. coli RNA polymerase with a bound fluoride riboswitch in the unliganded state type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#7 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 356 KDa |
-Macromolecule #1: DNA (39-MER)
| Macromolecule | Name: DNA (39-MER) / type: dna / ID: 1 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 7.942782 KDa |
| Sequence | String: (DG)(DT)(DC)(DC)(N)(N)(N)(N)(N)(N) (N)(N)(N)(N)(N)(N)(DG)(DA)(DA)(DG) (DA)(DG)(DA) (DT)(DT)(DC)(DA)(DG)(DA) (DG) |
-Macromolecule #2: DNA (30-MER)
| Macromolecule | Name: DNA (30-MER) / type: dna / ID: 2 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 9.03883 KDa |
| Sequence | String: (DC)(DT)(DC)(DT)(DG)(DA)(DA)(DT)(DC)(DT) (DC)(DT)(DT)(DC)(DC)(DA)(DC)(DT)(DC)(DC) (DT)(DA)(DC)(DC)(DA)(DA)(DG)(DG)(DA) (DC) |
-Macromolecule #3: DNA-directed RNA polymerase subunit alpha
| Macromolecule | Name: DNA-directed RNA polymerase subunit alpha / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO / EC number: DNA-directed RNA polymerase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 25.568074 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: VTEFLKPRLV DIEQVSSTHA KVTLEPLERG FGHTLGNALR RILLSSMPGC AVTEVEIDGV LHEYSTKEGV QEDILEILLN LKGLAVRVQ GKDEVILTLN KSGIGPVTAA DITHDGDVEI VKPQHVICHL TDENASISMR IKVQRGRGYV PASTRIHSEE D ERPIGRLL ...String: VTEFLKPRLV DIEQVSSTHA KVTLEPLERG FGHTLGNALR RILLSSMPGC AVTEVEIDGV LHEYSTKEGV QEDILEILLN LKGLAVRVQ GKDEVILTLN KSGIGPVTAA DITHDGDVEI VKPQHVICHL TDENASISMR IKVQRGRGYV PASTRIHSEE D ERPIGRLL VDACYSPVER IAYNVEAARV EQRTDLDKLV IEMETNGTID PEEAIRRAAT ILAEQLEAFV DLE UniProtKB: DNA-directed RNA polymerase subunit alpha |
-Macromolecule #4: DNA-directed RNA polymerase subunit beta
| Macromolecule | Name: DNA-directed RNA polymerase subunit beta / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO / EC number: DNA-directed RNA polymerase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 150.560562 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: VYSYTEKKRI RKDFGKRPQV LDVPYLLSIQ LDSFQKFIEQ DPEGQYGLEA AFRSVFPIQS YSGNSELQYV SYRLGEPVFD VQECQIRGV TYSAPLRVKL RLVIYEREAP EGTVKDIKEQ EVYMGEIPLM TDNGTFVING TERVIVSQLH RSPGVFFDSD K GKTHSSGK ...String: VYSYTEKKRI RKDFGKRPQV LDVPYLLSIQ LDSFQKFIEQ DPEGQYGLEA AFRSVFPIQS YSGNSELQYV SYRLGEPVFD VQECQIRGV TYSAPLRVKL RLVIYEREAP EGTVKDIKEQ EVYMGEIPLM TDNGTFVING TERVIVSQLH RSPGVFFDSD K GKTHSSGK VLYNARIIPY RGSWLDFEFD PKDNLFVRID RRRKLPATII LRALNYTTEQ ILDLFFEKVI FEIRDNKLQM EL VPERLRG ETASFDIEAN GKVYVEKGRR ITARHIRQLE KDDVKLIEVP VEYIAGKVVA KDYIDESTGE LICAANMELS LDL LAKLSQ SGHKRIETLF TNDLDHGPYI SETLRVDPTN DRLSALVEIY RMMRPGEPPT REAAESLFEN LFFSEDRYDL SAVG RMKFN RSLLREEIEG SGILSKDDII DVMKKLIDIR NGKGEVDDID HLGNRRIRSV GEMAENQFRV GLVRVERAVK ERLSL GDLD TLMPQDMINA KPISAAVKEF FGSSQLSQFM DQNNPLSEIT HKRRISALGP GGLTRERAGF EVRDVHPTHY GRVCPI ETP EGPNIGLINS LSVYAQTNEY GFLETPYRKV TDGVVTDEIH YLSAIEEGNY VIAQANSNLD EEGHFVEDLV TCRSKGE SS LFSRDQVDYM DVSTQQVVSV GASLIPFLEH DDANRALMGA NMQRQAVPTL RADKPLVGTG MERAVAVDSG VTAVAKRG G VVQYVDASRI VIKVNEDEMY PGEAGIDIYN LTKYTRSNQN TCINQMPCVS LGEPVERGDV LADGPSTDLG ELALGQNMR VAFMPWNGYN FEDSILVSER VVQEDRFTTI HIQELACVSR DTKLGPEEIT ADIPNVGEAA LSKLDESGIV YIGAEVTGGD ILVGKVTPK GETQLTPEEK LLRAIFGEKA SDVKDSSLRV PNGVSGTVID VQVFTRDGVE KDKRALEIEE MQLKQAKKDL S EELQILEA GLFSRIRAVL VAGGVEAEKL DKLPRDRWLE LGLTDEEKQN QLEQLAEQYD ELKHEFEKKL EAKRRKITQG DD LAPGVLK IVKVYLAVKR RIQPGDKMAG RHGNKGVISK INPIEDMPYD ENGTPVDIVL NPLGVPSRMN IGQILETHLG MAA KGIGDK INAMLKQQQE VAKLREFIQR AYDLGADVRQ KVDLSTFSDE EVMRLAENLR KGMPIATPVF DGAKEAEIKE LLKL GDLPT SGQIRLYDGR TGEQFERPVT VGYMYMLKLN HLVDDKMHAR STGSYSLVTQ QPLGGKAQFG GQRFGEMEVW ALEAY GAAY TLQEMLTVKS DDVNGRTKMY KNIVDGNHQM EPGMPESFNV LLKEIRSLGI NIELED UniProtKB: DNA-directed RNA polymerase subunit beta |
-Macromolecule #5: DNA-directed RNA polymerase subunit beta'
| Macromolecule | Name: DNA-directed RNA polymerase subunit beta' / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO / EC number: DNA-directed RNA polymerase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 150.436344 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: EFDAIKIALA SPDMIRSWSF GEVKKPETIN YRTFKPERDG LFCARIFGPV KDYECLCGKY KRLKHRGVIC EKCGVEVTQT KVRRERMGH IELASPTAHI WFLKSLPSRI GLLLDMPLRD IERVLYFESY VVIEGGMTNL ERQQILTEEQ YLDALEEFGD E FDAKMGAE ...String: EFDAIKIALA SPDMIRSWSF GEVKKPETIN YRTFKPERDG LFCARIFGPV KDYECLCGKY KRLKHRGVIC EKCGVEVTQT KVRRERMGH IELASPTAHI WFLKSLPSRI GLLLDMPLRD IERVLYFESY VVIEGGMTNL ERQQILTEEQ YLDALEEFGD E FDAKMGAE AIQALLKSMD LEQECEQLRE ELNETNSETK RKKLTKRIKL LEAFVQSGNK PEWMILTVLP VLPPDLRPLV PL DGGRFAT SDLNDLYRRV INRNNRLKRL LDLAAPDIIV RNEKRMLQEA VDALLDNGRR GRAITGSNKR PLKSLADMIK GKQ GRFRQN LLGKRVDYSG RSVITVGPYL RLHQCGLPKK MALELFKPFI YGKLELRGLA TTIKAAKKMV EREEAVVWDI LDEV IREHP VLLNRAPTLH RLGIQAFEPV LIEGKAIQLH PLVCAAYNAD FDGDQMAVHV PLTLEAQLEA RALMMSTNNI LSPAN GEPI IVPSQDVVLG LYYMTRDCVN AKGEGMVLTG PKEAERLYRS GLASLHARVK VRITEYEKDA NGELVAKTSL KDTTVG RAI LWMIVPKGLP YSIVNQALGK KAISKMLNTC YRILGLKPTV IFADQIMYTG FAYAARSGAS VGIDDMVIPE KKHEIIS EA EAEVAEIQEQ FQSGLVTAGE RYNKVIDIWA AANDRVSKAM MDNLQTETVI NRDGQEEKQV SFNSIYMMAD SGARGSAA Q IRQLAGMRGL MAKPDGSIIE TPITANFREG LNVLQYFIST HGARKGLADT ALKTANSGYL TRRLVDVAQD LVVTEDDCG THEGIMMTPV IEGGDVKEPL RDRVLGRVTA EDVLKPGTAD ILVPRNTLLH EQWCDLLEEN SVDAVKVRSV VSCDTDFGVC AHCYGRDLA RGHIINKGEA IGVIAAQSIG EPGTQLTMRT FHIGGAASRA AAESSIQVKN KGSIKLSNVK SVVNSSGKLV I TSRNTELK LIDEFGRTKE SYKVPYGAVL AKGDGEQVAG GETVANWDPH TMPVITEVSG FVRFTDMIDG QTITRQTDEL TG LSSLVVL DSAERTAGGK DLRPALKIVD AQGNDVLIPG TDMPAQYFLP GKAIVQLEDG VQISSGDTLA RIPQESGGTK DIT GGLPRV ADLFEARRPK EPAILAEISG IVSFGKETKG KRRLVITPVD GSDPYEEMIP KWRQLNVFEG ERVERGDVIS DGPE APHDI LRLRGVHAVT RYIVNEVQDV YRLQGVKIND KHIEVIVRQM LRKATIVNAG SSDFLEGEQV EYSRVKIANR ELEAN GKVG ATYSRDLLGI TKASLATESF ISAASFQETT RVLTEAAVAG KRDELRGLKE NVIVGRLIPA GTGYAYHQDR MRRR UniProtKB: DNA-directed RNA polymerase subunit beta' |
-Macromolecule #6: DNA-directed RNA polymerase subunit omega
| Macromolecule | Name: DNA-directed RNA polymerase subunit omega / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO / EC number: DNA-directed RNA polymerase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 8.963044 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: ARVTVQDAVE KIGNRFDLVL VAARRARQMQ VGGKDPLVPE ENDKTTVIAL REIEEGLINN QILDVRERQE QQEQEAAEL UniProtKB: DNA-directed RNA polymerase subunit omega |
-Macromolecule #7: RNA (5'-R(P*AP*GP*UP*AP*UP*CP*AP*CP*UP*AP*CP*UP*GP*GP*UP*AP*GP*GP...
| Macromolecule | Name: RNA (5'-R(P*AP*GP*UP*AP*UP*CP*AP*CP*UP*AP*CP*UP*GP*GP*UP*AP*GP*GP*AP*GP*U)-3') type: rna / ID: 7 / Number of copies: 1 / EC number: DNA-directed RNA polymerase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 6.754054 KDa |
| Sequence | String: AGUAUCACUA CUGGUAGGAG U |
-Macromolecule #8: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 8 / Number of copies: 1 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: 5 mM Tris-HCl, pH 7.5, 100 mM KCl, 1 mM MgCl2 | ||||||||||||
| Grid | Model: C-flat / Material: GOLD / Mesh: 400 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 52.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)