Entry Database : EMDB / ID : EMD-45805Title Infectious bronchitis virus core polymerase complex Complex : Core polymerase complex of the infectious bronchitis virus composed of nsp12, nsp8, nsp7, and an dsRNA substrateProtein or peptide : RNA-directed RNA polymerase nsp12Protein or peptide : Non-structural protein 8Protein or peptide : Non-structural protein 7RNA : RNA PrimerRNA : RNA templateLigand : ZINC ION / / / / / Function / homology Function Domain/homology Component
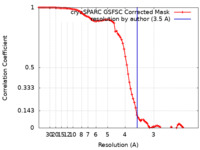
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species / Method / / Resolution : 3.5 Å Hoferle PJ / Anderson TK / Kirchdoerfer RN Funding support Organization Grant number Country National Institutes of Health/National Institute Of Allergy and Infectious Diseases (NIH/NIAID) AI158463
Journal : Biorxiv / Year : 2024Title : A genus-specific nsp12 region impacts polymerase assembly in Alpha- and GammacoronavirusesAuthors : Hoferle PJ / Anderson TK / Kirchdoerfer RN History Deposition Jul 18, 2024 - Header (metadata) release Jul 31, 2024 - Map release Jul 31, 2024 - Update Jul 31, 2024 - Current status Jul 31, 2024 Processing site : RCSB / Status : Released
Show all Show less
 Open data
Open data Basic information
Basic information
 Map data
Map data Sample
Sample Keywords
Keywords Function and homology information
Function and homology information Infectious bronchitis virus /
Infectious bronchitis virus / 
 Authors
Authors United States, 1 items
United States, 1 items  Citation
Citation Journal: Biorxiv / Year: 2024
Journal: Biorxiv / Year: 2024 Structure visualization
Structure visualization Downloads & links
Downloads & links emd_45805.map.gz
emd_45805.map.gz EMDB map data format
EMDB map data format emd-45805-v30.xml
emd-45805-v30.xml emd-45805.xml
emd-45805.xml EMDB header
EMDB header emd_45805_fsc.xml
emd_45805_fsc.xml FSC data file
FSC data file emd_45805.png
emd_45805.png emd-45805.cif.gz
emd-45805.cif.gz emd_45805_half_map_1.map.gz
emd_45805_half_map_1.map.gz emd_45805_half_map_2.map.gz
emd_45805_half_map_2.map.gz http://ftp.pdbj.org/pub/emdb/structures/EMD-45805
http://ftp.pdbj.org/pub/emdb/structures/EMD-45805 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-45805
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-45805 emd_45805_validation.pdf.gz
emd_45805_validation.pdf.gz EMDB validaton report
EMDB validaton report emd_45805_full_validation.pdf.gz
emd_45805_full_validation.pdf.gz emd_45805_validation.xml.gz
emd_45805_validation.xml.gz emd_45805_validation.cif.gz
emd_45805_validation.cif.gz https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-45805
https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-45805 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-45805
ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-45805
 F&H Search
F&H Search Links
Links EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource Map
Map Download / File: emd_45805.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
Download / File: emd_45805.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Sample components
Sample components Infectious bronchitis virus
Infectious bronchitis virus Infectious bronchitis virus
Infectious bronchitis virus
 Infectious bronchitis virus
Infectious bronchitis virus
 Infectious bronchitis virus
Infectious bronchitis virus


 Processing
Processing Sample preparation
Sample preparation Electron microscopy
Electron microscopy FIELD EMISSION GUN
FIELD EMISSION GUN
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)