+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CCHFV GP38 bound to ADI-46152 and ADI-58048 Fabs | |||||||||
 Map data Map data | Sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CCHFV / GP38 / Antibody / Immunology / VIRAL PROTEIN / VIRAL PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationhost cell Golgi membrane / clathrin-dependent endocytosis of virus by host cell / host cell endoplasmic reticulum membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / virion membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  Crimean-Congo hemorrhagic fever virus / Crimean-Congo hemorrhagic fever virus /  Crimean-Congo hemorrhagic fever virus strain IbAr10200 Crimean-Congo hemorrhagic fever virus strain IbAr10200 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.9 Å | |||||||||
 Authors Authors | Hjorth CK / McLellan JS | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2024 Journal: Cell Rep / Year: 2024Title: Crimean-Congo hemorrhagic fever survivors elicit protective non-neutralizing antibodies that target 11 overlapping regions on glycoprotein GP38. Authors: Olivia S Shin / Stephanie R Monticelli / Christy K Hjorth / Vladlena Hornet / Michael Doyle / Dafna Abelson / Ana I Kuehne / Albert Wang / Russell R Bakken / Akaash K Mishra / Marissa ...Authors: Olivia S Shin / Stephanie R Monticelli / Christy K Hjorth / Vladlena Hornet / Michael Doyle / Dafna Abelson / Ana I Kuehne / Albert Wang / Russell R Bakken / Akaash K Mishra / Marissa Middlecamp / Elizabeth Champney / Lauran Stuart / Daniel P Maurer / Jiannan Li / Jacob Berrigan / Jennifer Barajas / Stephen Balinandi / Julius J Lutwama / Leslie Lobel / Larry Zeitlin / Laura M Walker / John M Dye / Kartik Chandran / Andrew S Herbert / Noel T Pauli / Jason S McLellan /   Abstract: Crimean-Congo hemorrhagic fever virus can cause lethal disease in humans yet there are no approved medical countermeasures. Viral glycoprotein GP38, exclusive to Nairoviridae, is a target of ...Crimean-Congo hemorrhagic fever virus can cause lethal disease in humans yet there are no approved medical countermeasures. Viral glycoprotein GP38, exclusive to Nairoviridae, is a target of protective antibodies and is a key antigen in preclinical vaccine candidates. Here, we isolate 188 GP38-specific antibodies from human survivors of infection. Competition experiments show that these antibodies bind across 5 distinct antigenic sites, encompassing 11 overlapping regions. Additionally, we show structures of GP38 bound with 9 of these antibodies targeting different antigenic sites. Although these GP38-specific antibodies are non-neutralizing, several display protective efficacy equal to or better than murine antibody 13G8 in two highly stringent rodent models of infection. Together, these data expand our understanding regarding this important viral protein and may inform the development of broadly effective CCHFV antibody therapeutics. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43604.map.gz emd_43604.map.gz | 259.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43604-v30.xml emd-43604-v30.xml emd-43604.xml emd-43604.xml | 23.9 KB 23.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_43604_fsc.xml emd_43604_fsc.xml | 13.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_43604.png emd_43604.png | 53.4 KB | ||
| Filedesc metadata |  emd-43604.cif.gz emd-43604.cif.gz | 6.6 KB | ||
| Others |  emd_43604_half_map_1.map.gz emd_43604_half_map_1.map.gz emd_43604_half_map_2.map.gz emd_43604_half_map_2.map.gz | 254.9 MB 254.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43604 http://ftp.pdbj.org/pub/emdb/structures/EMD-43604 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43604 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43604 | HTTPS FTP |
-Related structure data
| Related structure data |  8vwwMC  8vvkC  8vvlC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_43604.map.gz / Format: CCP4 / Size: 274.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43604.map.gz / Format: CCP4 / Size: 274.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8332 Å | ||||||||||||||||||||||||||||||||||||
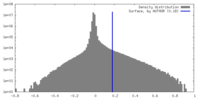
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half map A
| File | emd_43604_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
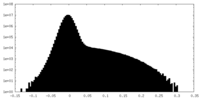
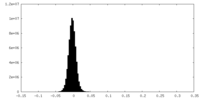
| Density Histograms |
-Half map: Half map B
| File | emd_43604_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
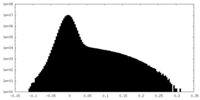
| Density Histograms |
- Sample components
Sample components
-Entire : CCHFV GP38 bound with ADI-46152 and ADI-58048 Fabs
| Entire | Name: CCHFV GP38 bound with ADI-46152 and ADI-58048 Fabs |
|---|---|
| Components |
|
-Supramolecule #1: CCHFV GP38 bound with ADI-46152 and ADI-58048 Fabs
| Supramolecule | Name: CCHFV GP38 bound with ADI-46152 and ADI-58048 Fabs / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Supramolecule #2: ADI-46152 Fab
| Supramolecule | Name: ADI-46152 Fab / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: CCHFV GP38
| Supramolecule | Name: CCHFV GP38 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Crimean-Congo hemorrhagic fever virus / Strain: IbAr10200 Crimean-Congo hemorrhagic fever virus / Strain: IbAr10200 |
-Supramolecule #4: ADI-58048 Fab
| Supramolecule | Name: ADI-58048 Fab / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #4-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: GP38
| Macromolecule | Name: GP38 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Crimean-Congo hemorrhagic fever virus strain IbAr10200 Crimean-Congo hemorrhagic fever virus strain IbAr10200Strain: IbAr10200 |
| Molecular weight | Theoretical: 30.241828 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: NLKMEIILTL SQGLKKYYGK ILRLLQLTLE EDTEGLLEWC KRNLGLDCDD TFFQKRIEEF FITGEGHFNE VLQFRTPGTL STTESTPAG LPTAEPFKSY FAKGFLSIDS GYYSAKCYSG TSNSGLQLIN ITRHSTRIVD TPGPKITNLK TINCINLKAS I FKEHREVE ...String: NLKMEIILTL SQGLKKYYGK ILRLLQLTLE EDTEGLLEWC KRNLGLDCDD TFFQKRIEEF FITGEGHFNE VLQFRTPGTL STTESTPAG LPTAEPFKSY FAKGFLSIDS GYYSAKCYSG TSNSGLQLIN ITRHSTRIVD TPGPKITNLK TINCINLKAS I FKEHREVE INVLLPQVAV NLSNCHVVIK SHVCDYSLDI DGAVRLPHIY HEGVFIPGTY KIVIDKKNKL NDRCTLFTDC VI KGREVRK GQSVLRQYKT EIRIGKASTG S UniProtKB: Envelopment polyprotein |
-Macromolecule #2: ADI-46152 Fab Heavy Chain
| Macromolecule | Name: ADI-46152 Fab Heavy Chain / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 24.370311 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EVQLVESGGV LVQPGGSLRL SCAASGFTVN SNYMTWVRQA PGKGLEWVSV IYSGGYTYYA DSVKGRFAIS RDNSKNTVYL QMNSLRVED TAVYYCARLR LSSSWYPEAF DYWGQGTLVT VSSASTKGPS VFPLAPSSKS TSGGTAALGC LVKDYFPEPV T VSWNSGAL ...String: EVQLVESGGV LVQPGGSLRL SCAASGFTVN SNYMTWVRQA PGKGLEWVSV IYSGGYTYYA DSVKGRFAIS RDNSKNTVYL QMNSLRVED TAVYYCARLR LSSSWYPEAF DYWGQGTLVT VSSASTKGPS VFPLAPSSKS TSGGTAALGC LVKDYFPEPV T VSWNSGAL TSGVHTFPAV LQSSGLYSLS SVVTVPSSSL GTQTYICNVN HKPSNTKVDK KVEPKSCDKG |
-Macromolecule #3: ADI-46152 Fab Light Chain
| Macromolecule | Name: ADI-46152 Fab Light Chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 22.957463 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AILLTQSPSS LSASVGDRVT ITCRASQGIS SALAWYQQKP GRAPKVLIYD ASSLANGVPS RFSGSGSGTD FTLTINSLQP EDFATYYCQ QFNYYPLTFG GGTKVEIKRT VAAPSVFIFP PSDEQLKSGT ASVVCLLNNF YPREAKVQWK VDNALQSGNS Q ESVTEQDS ...String: AILLTQSPSS LSASVGDRVT ITCRASQGIS SALAWYQQKP GRAPKVLIYD ASSLANGVPS RFSGSGSGTD FTLTINSLQP EDFATYYCQ QFNYYPLTFG GGTKVEIKRT VAAPSVFIFP PSDEQLKSGT ASVVCLLNNF YPREAKVQWK VDNALQSGNS Q ESVTEQDS KDSTYSLSST LTLSKADYEK HKVYACEVTQ GTTSVTKSFN RGEC |
-Macromolecule #4: ADI-58048 Fab Heavy Chain
| Macromolecule | Name: ADI-58048 Fab Heavy Chain / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.790645 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QLQLQESGPG LVKPSETLSL TCTVSGGSIT TSHYYWGWIR QPPGKGLEWV GSMYYSGGIY YNPSLKGRVT ISVDTSKNQF SLKLSSVTA ADTAVYYCAL ADAPDDAFDI WGQGTMVTVS SASTKGPSVF PLAPSSKSTS GGTAALGCLV KDYFPEPVTV S WNSGALTS ...String: QLQLQESGPG LVKPSETLSL TCTVSGGSIT TSHYYWGWIR QPPGKGLEWV GSMYYSGGIY YNPSLKGRVT ISVDTSKNQF SLKLSSVTA ADTAVYYCAL ADAPDDAFDI WGQGTMVTVS SASTKGPSVF PLAPSSKSTS GGTAALGCLV KDYFPEPVTV S WNSGALTS GVHTFPAVLQ SSGLYSLSSV VTVPSSSLGT QTYICNVNHK PSNTKVDKKV EPKSCDKG |
-Macromolecule #5: ADI-58048 Fab Light Chain
| Macromolecule | Name: ADI-58048 Fab Light Chain / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.36591 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DIQMTQSPSS LSAFVGDRVT ITCRASQSIS SYLNWYQQKP GKAPKLLIYA ASTLQSGVPS RFSGSGSGTD FTLTISSLQS EDFATYYCQ ESYSIPFTFG PGTKVDIKRT VAAPSVFIFP PSDEQLKSGT ASVVCLLNNF YPREAKVQWK VDNALQSGNS Q ESVTEQDS ...String: DIQMTQSPSS LSAFVGDRVT ITCRASQSIS SYLNWYQQKP GKAPKLLIYA ASTLQSGVPS RFSGSGSGTD FTLTISSLQS EDFATYYCQ ESYSIPFTFG PGTKVDIKRT VAAPSVFIFP PSDEQLKSGT ASVVCLLNNF YPREAKVQWK VDNALQSGNS Q ESVTEQDS KDSTYSLSST LTLSKADYEK HKVYACEVTH QGLSSPVTKS FNRGEC |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 3647 / Average electron dose: 80.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 105000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT | ||||||||||||
| Output model |  PDB-8vww: |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)