[English] 日本語
 Yorodumi
Yorodumi- EMDB-43190: HIV Env BG505_MD39_B16 SOSIP boosting trimer in complex with B16_... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | HIV Env BG505_MD39_B16 SOSIP boosting trimer in complex with B16_d77.5 mouse Fab and RM20A3 Fab | |||||||||
 Map data Map data | sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | mouse antibody / germline targeting / HIV-1 / vaccine design / VIRAL PROTEIN / VIRAL PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell ...symbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / host cell plasma membrane / virion membrane / structural molecule activity / membrane / identical protein binding Similarity search - Function | |||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /   | |||||||||
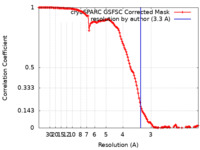
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Ozorowski G / Torres JL / Ward AB | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2024 Journal: Science / Year: 2024Title: mRNA-LNP HIV-1 trimer boosters elicit precursors to broad neutralizing antibodies. Authors: Zhenfei Xie / Ying-Cing Lin / Jon M Steichen / Gabriel Ozorowski / Sven Kratochvil / Rashmi Ray / Jonathan L Torres / Alessia Liguori / Oleksandr Kalyuzhniy / Xuesong Wang / John E Warner / ...Authors: Zhenfei Xie / Ying-Cing Lin / Jon M Steichen / Gabriel Ozorowski / Sven Kratochvil / Rashmi Ray / Jonathan L Torres / Alessia Liguori / Oleksandr Kalyuzhniy / Xuesong Wang / John E Warner / Stephanie R Weldon / Gordon A Dale / Kathrin H Kirsch / Usha Nair / Sabyasachi Baboo / Erik Georgeson / Yumiko Adachi / Michael Kubitz / Abigail M Jackson / Sara T Richey / Reid M Volk / Jeong Hyun Lee / Jolene K Diedrich / Thavaleak Prum / Samantha Falcone / Sunny Himansu / Andrea Carfi / John R Yates / James C Paulson / Devin Sok / Andrew B Ward / William R Schief / Facundo D Batista /  Abstract: Germline-targeting (GT) HIV vaccine strategies are predicated on deriving broadly neutralizing antibodies (bnAbs) through multiple boost immunogens. However, as the recruitment of memory B cells ...Germline-targeting (GT) HIV vaccine strategies are predicated on deriving broadly neutralizing antibodies (bnAbs) through multiple boost immunogens. However, as the recruitment of memory B cells (MBCs) to germinal centers (GCs) is inefficient and may be derailed by serum antibody-induced epitope masking, driving further B cell receptor (BCR) modification in GC-experienced B cells after boosting poses a challenge. Using humanized immunoglobulin knockin mice, we found that GT protein trimer immunogen N332-GT5 could prime inferred-germline precursors to the V3-glycan-targeted bnAb BG18 and that B cells primed by N332-GT5 were effectively boosted by either of two novel protein immunogens designed to have minimum cross-reactivity with the off-target V1-binding responses. The delivery of the prime and boost immunogens as messenger RNA lipid nanoparticles (mRNA-LNPs) generated long-lasting GCs, somatic hypermutation, and affinity maturation and may be an effective tool in HIV vaccine development. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43190.map.gz emd_43190.map.gz | 230.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43190-v30.xml emd-43190-v30.xml emd-43190.xml emd-43190.xml | 28.4 KB 28.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_43190_fsc.xml emd_43190_fsc.xml | 13.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_43190.png emd_43190.png | 98.8 KB | ||
| Masks |  emd_43190_msk_1.map emd_43190_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-43190.cif.gz emd-43190.cif.gz | 8.4 KB | ||
| Others |  emd_43190_half_map_1.map.gz emd_43190_half_map_1.map.gz emd_43190_half_map_2.map.gz emd_43190_half_map_2.map.gz | 226.7 MB 226.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43190 http://ftp.pdbj.org/pub/emdb/structures/EMD-43190 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43190 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43190 | HTTPS FTP |
-Related structure data
| Related structure data |  8vfvMC  8f92C  8f9gC  8f9mC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_43190.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43190.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.044 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_43190_msk_1.map emd_43190_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map A
| File | emd_43190_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map B
| File | emd_43190_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : IV Env BG505_MD39_B16 SOSIP boosting trimer in complex with B16_d...
| Entire | Name: IV Env BG505_MD39_B16 SOSIP boosting trimer in complex with B16_d77.5 mouse Fab and RM20A3 Fab |
|---|---|
| Components |
|
-Supramolecule #1: IV Env BG505_MD39_B16 SOSIP boosting trimer in complex with B16_d...
| Supramolecule | Name: IV Env BG505_MD39_B16 SOSIP boosting trimer in complex with B16_d77.5 mouse Fab and RM20A3 Fab type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
-Macromolecule #1: RM20A3 Fab heavy chain
| Macromolecule | Name: RM20A3 Fab heavy chain / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.511111 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EVQLVETGGG LVQPGGSLKL SCRASGYTFS SFAMSWVRQA PGKGLEWVSL INDRGGLTFY VDSVKGRFTI SRDNSKNTLS LQMHSLRDG DTAVYYCATG GMSSALQSSK YYFDFWGQGA LVTVSS |
-Macromolecule #2: RM20A3 Fab light chain
| Macromolecule | Name: RM20A3 Fab light chain / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.5088 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: ALTQPPSVSG SPGQSVTISC TGTSSDIGSY NYVSWYQQHP GKAPKLMIYD VTQRPSGVSD RFSGSKSGNT ASLTISGLQA DDEADYYCS AYAGRQTFYI FGGGTRLTVL GQPKASPTVT LFPPSSEEL |
-Macromolecule #3: Envelope glycoprotein gp120
| Macromolecule | Name: Envelope glycoprotein gp120 / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 54.221656 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AENLWVTVYY GVPVWKDAET TLFCASDAKA YETEKHNVWA THACVPTDPN PQEIHLENVT EEFNMWKNNM VEQMHEDIIS LWDQSLKPC VKLTPLCVTL QCTNYTEKLR SMMKGELKNC SFNMTTELRD KKQKVYSLFY RLDVVQINEN QGNRSNNSNK E YRLINCNT ...String: AENLWVTVYY GVPVWKDAET TLFCASDAKA YETEKHNVWA THACVPTDPN PQEIHLENVT EEFNMWKNNM VEQMHEDIIS LWDQSLKPC VKLTPLCVTL QCTNYTEKLR SMMKGELKNC SFNMTTELRD KKQKVYSLFY RLDVVQINEN QGNRSNNSNK E YRLINCNT SAITQACPKV SFEPIPIHYC APAGFAILKC KDKKFNGTGP CPSVSTVQCT HGIKPVVSTQ LLLNGSLAEE EV IIRSENI TNNAKNILVQ LNTPVQINCT RPNNNTVKSI RIGPGQAFYY TGDIIGHIRQ AHCNVSKATW NETLGKVVKQ LRK HFGNNT IIRFAQSSGG DLEVTTHSFN CGGEFFYCNT SGLFNSTWIS NTSVQGSNST GSNDSITLPC RIKQIINMWQ RIGQ AMYAP PIQGVIRCVS NITGLILTRD GGSTNSTTET FRPGGGDMRD NWRSELYKYK VVKIEPLGVA PTRCKRRVVG RRRRR R UniProtKB: Envelope glycoprotein gp160 |
-Macromolecule #4: Transmembrane protein gp41
| Macromolecule | Name: Transmembrane protein gp41 / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 17.134324 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AVGIGAVSLG FLGAAGSTMG AASMTLTVQA RNLLSGIVQQ QSNLLRAPEP QQHLLKDTHW GIKQLQARVL AVEHYLRDQQ LLGIWGCSG KLICCTNVPW NSSWSNRNLS EIWDNMTWLQ WDKEISNYTQ IIYGLLEESQ NQQEKNEQDL LALD UniProtKB: Envelope glycoprotein gp160 |
-Macromolecule #5: B16_d77.5 mouse Fab heavy chain Fv
| Macromolecule | Name: B16_d77.5 mouse Fab heavy chain Fv / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.972703 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLQESGPG LVKPSETLSL TCAVSGGSIS SGHWWSWVRQ SPGKGLEWIG TIYHSGSANY NPSLKSRVTI SVDKSKNQFS LKLTSVTAA DTAVYFCARN AILIFGVIAF GEYYFCGMDV WGQGTTVTVS S |
-Macromolecule #6: B16_d77.5 mouse Fab light chain Fv
| Macromolecule | Name: B16_d77.5 mouse Fab light chain Fv / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 12.012308 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DIVLTQSPAS LAVSLGQRAT IFCRASDTVD ISGDSFMHWY QQKPGQPPNL LIYRASNLQS GIPARFSGSG SRADFTLTID PVEADDVAT YYCQQSSEDP LTFGAGTKLE LK |
-Macromolecule #10: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 10 / Number of copies: 36 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 6.9 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: TFS FALCON 4i (4k x 4k) / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 190000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)