[English] 日本語
 Yorodumi
Yorodumi- EMDB-41631: Cryo-EM structure of Chikungunya virus with asymmetric reconstruction -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Chikungunya virus with asymmetric reconstruction | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Chikungunya virus / VIRUS | |||||||||
| Biological species |   Chikungunya virus Chikungunya virus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 6.6 Å | |||||||||
 Authors Authors | Su GC / Chmielewsk D / Kaelber J / Pintilie G / Chen M / Jin J / Auguste A / Chiu W | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: PNAS Nexus / Year: 2024 Journal: PNAS Nexus / Year: 2024Title: Cryogenic electron microscopy and tomography reveal imperfect icosahedral symmetry in alphaviruses. Authors: David Chmielewski / Guan-Chin Su / Jason T Kaelber / Grigore D Pintilie / Muyuan Chen / Jing Jin / Albert J Auguste / Wah Chiu /  Abstract: Alphaviruses are spherical, enveloped RNA viruses with two-layered icosahedral architecture. The structures of many alphaviruses have been studied using cryogenic electron microscopy (cryo-EM) ...Alphaviruses are spherical, enveloped RNA viruses with two-layered icosahedral architecture. The structures of many alphaviruses have been studied using cryogenic electron microscopy (cryo-EM) reconstructions, which impose icosahedral symmetry on the viral particles. Using cryogenic electron tomography (cryo-ET), we revealed a polarized symmetry defect in the icosahedral lattice of Chikungunya virus (CHIKV) in situ, similar to the late budding particles, suggesting the inherent imperfect symmetry originates from the final pinch-off of assembled virions. We further demonstrated this imperfect symmetry is also present in in vitro purified CHIKV and Mayaro virus, another arthritogenic alphavirus. We employed a subparticle-based single-particle analysis protocol to circumvent the icosahedral imperfection and boosted the resolution of the structure of the CHIKV to ∼3 Å resolution, which revealed detailed molecular interactions between glycoprotein E1-E2 heterodimers in the transmembrane region and multiple lipid-like pocket factors located in a highly conserved hydrophobic pocket. This complementary use of in situ cryo-ET and single-particle cryo-EM approaches provides a more precise structural description of near-icosahedral viruses and valuable insights to guide the development of structure-based antiviral therapies against alphaviruses. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41631.map.gz emd_41631.map.gz | 47.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41631-v30.xml emd-41631-v30.xml emd-41631.xml emd-41631.xml | 15.4 KB 15.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_41631_fsc.xml emd_41631_fsc.xml | 12.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_41631.png emd_41631.png | 109.4 KB | ||
| Filedesc metadata |  emd-41631.cif.gz emd-41631.cif.gz | 4.5 KB | ||
| Others |  emd_41631_half_map_1.map.gz emd_41631_half_map_1.map.gz emd_41631_half_map_2.map.gz emd_41631_half_map_2.map.gz | 129.1 MB 128.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41631 http://ftp.pdbj.org/pub/emdb/structures/EMD-41631 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41631 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41631 | HTTPS FTP |
-Validation report
| Summary document |  emd_41631_validation.pdf.gz emd_41631_validation.pdf.gz | 1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_41631_full_validation.pdf.gz emd_41631_full_validation.pdf.gz | 1 MB | Display | |
| Data in XML |  emd_41631_validation.xml.gz emd_41631_validation.xml.gz | 19.7 KB | Display | |
| Data in CIF |  emd_41631_validation.cif.gz emd_41631_validation.cif.gz | 26.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41631 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41631 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41631 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41631 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_41631.map.gz / Format: CCP4 / Size: 163.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41631.map.gz / Format: CCP4 / Size: 163.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.68 Å | ||||||||||||||||||||||||||||||||||||
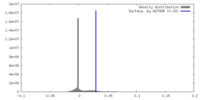
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_41631_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
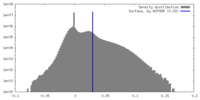
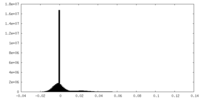
| Density Histograms |
-Half map: #1
| File | emd_41631_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
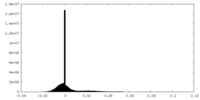
| Density Histograms |
- Sample components
Sample components
-Entire : Chikungunya virus
| Entire | Name:   Chikungunya virus Chikungunya virus |
|---|---|
| Components |
|
-Supramolecule #1: Chikungunya virus
| Supramolecule | Name: Chikungunya virus / type: virus / ID: 1 / Parent: 0 / NCBI-ID: 37124 / Sci species name: Chikungunya virus / Sci species strain: vaccine strain 181/clone 25 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No |
|---|---|
| Molecular weight | Theoretical: 500 KDa |
| Virus shell | Shell ID: 1 / Diameter: 700.0 Å / T number (triangulation number): 4 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293.15 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 47.25 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 106000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)