[English] 日本語
 Yorodumi
Yorodumi- EMDB-41440: Cryo-EM structure of TRNM-f*01 Fab in complex with HIV-1 Env trim... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of TRNM-f*01 Fab in complex with HIV-1 Env trimer ConC SOSIP | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Antibody / Vaccination / VIRAL PROTEIN / IMMUNE SYSTEM / VIRAL PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.03 Å | |||||||||
 Authors Authors | Roark RS / Morano NC / Shapiro LS / Kwong PD | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2024 Journal: Cell / Year: 2024Title: Potent and broad HIV-1 neutralization in fusion peptide-primed SHIV-infected macaques. Authors: Hua Wang / Cheng Cheng / James L Dal Santo / Chen-Hsiang Shen / Tatsiana Bylund / Amy R Henry / Colin A Howe / Juyun Hwang / Nicholas C Morano / Daniel J Morris / Sergei Pletnev / Ryan S ...Authors: Hua Wang / Cheng Cheng / James L Dal Santo / Chen-Hsiang Shen / Tatsiana Bylund / Amy R Henry / Colin A Howe / Juyun Hwang / Nicholas C Morano / Daniel J Morris / Sergei Pletnev / Ryan S Roark / Tongqing Zhou / Bryan T Hansen / Forrest H Hoyt / Timothy S Johnston / Shuyi Wang / Baoshan Zhang / David R Ambrozak / Jordan E Becker / Michael F Bender / Anita Changela / Ridhi Chaudhary / Martin Corcoran / Angela R Corrigan / Kathryn E Foulds / Yicheng Guo / Myungjin Lee / Yingying Li / Bob C Lin / Tracy Liu / Mark K Louder / Marco Mandolesi / Rosemarie D Mason / Krisha McKee / Vinod Nair / Sijy O'Dell / Adam S Olia / Li Ou / Amarendra Pegu / Nagarajan Raju / Reda Rawi / Jesmine Roberts-Torres / Edward K Sarfo / Mallika Sastry / Andrew J Schaub / Stephen D Schmidt / Chaim A Schramm / Cindi L Schwartz / Sarah C Smith / Tyler Stephens / Jonathan Stuckey / I-Ting Teng / John-Paul Todd / Yaroslav Tsybovsky / David J Van Wazer / Shuishu Wang / Nicole A Doria-Rose / Elizabeth R Fischer / Ivelin S Georgiev / Gunilla B Karlsson Hedestam / Zizhang Sheng / Ruth A Woodward / Daniel C Douek / Richard A Koup / Theodore C Pierson / Lawrence Shapiro / George M Shaw / John R Mascola / Peter D Kwong /   Abstract: An antibody-based HIV-1 vaccine will require the induction of potent cross-reactive HIV-1-neutralizing responses. To demonstrate feasibility toward this goal, we combined vaccination targeting the ...An antibody-based HIV-1 vaccine will require the induction of potent cross-reactive HIV-1-neutralizing responses. To demonstrate feasibility toward this goal, we combined vaccination targeting the fusion-peptide site of vulnerability with infection by simian-human immunodeficiency virus (SHIV). In four macaques with vaccine-induced neutralizing responses, SHIV infection boosted plasma neutralization to 45%-77% breadth (geometric mean 50% inhibitory dilution [ID] ∼100) on a 208-strain panel. Molecular dissection of these responses by antibody isolation and cryo-electron microscopy (cryo-EM) structure determination revealed 15 of 16 antibody lineages with cross-clade neutralization to be directed toward the fusion-peptide site of vulnerability. In each macaque, isolated antibodies from memory B cells recapitulated the plasma-neutralizing response, with fusion-peptide-binding antibodies reaching breadths of 40%-60% (50% inhibitory concentration [IC] < 50 μg/mL) and total lineage-concentrations estimates of 50-200 μg/mL. Longitudinal mapping indicated that these responses arose prior to SHIV infection. Collectively, these results provide in vivo molecular examples for one to a few B cell lineages affording potent, broadly neutralizing plasma responses. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41440.map.gz emd_41440.map.gz | 106.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41440-v30.xml emd-41440-v30.xml emd-41440.xml emd-41440.xml | 25.3 KB 25.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_41440.png emd_41440.png | 113.1 KB | ||
| Filedesc metadata |  emd-41440.cif.gz emd-41440.cif.gz | 7.6 KB | ||
| Others |  emd_41440_half_map_1.map.gz emd_41440_half_map_1.map.gz emd_41440_half_map_2.map.gz emd_41440_half_map_2.map.gz | 200.7 MB 200.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41440 http://ftp.pdbj.org/pub/emdb/structures/EMD-41440 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41440 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41440 | HTTPS FTP |
-Related structure data
| Related structure data |  8to9MC  8tdxC  8te7C  8tjrC  8tjsC  8tkcC  8tl2C  8tl3C  8tl4C  8tl5C  8tnuC  8to7C  8topC M: atomic model generated by this map C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_41440.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41440.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_41440_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
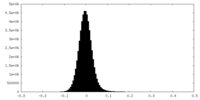
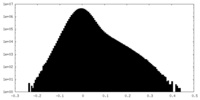
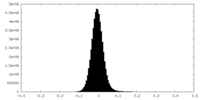
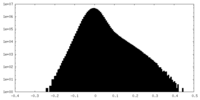
| Density Histograms |
-Half map: #1
| File | emd_41440_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : TRNM-f*01 Fab in complex with HIV-1 Env trimer ConC SOSIP
| Entire | Name: TRNM-f*01 Fab in complex with HIV-1 Env trimer ConC SOSIP |
|---|---|
| Components |
|
-Supramolecule #1: TRNM-f*01 Fab in complex with HIV-1 Env trimer ConC SOSIP
| Supramolecule | Name: TRNM-f*01 Fab in complex with HIV-1 Env trimer ConC SOSIP type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Envelope glycoprotein gp120
| Macromolecule | Name: Envelope glycoprotein gp120 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 52.737102 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: NLWVTVYYGV PVWKEAKTTL FCASDAKAYE KEVHNVWATH ACVPTDPNPQ EMVLENVTEN FNMWKNDMVD QMHEDIISLW DQSLKPCVK LTPLCVTLNC TNVNVTNTNN NNMKEEMKNC SFNTTTEIRD KKQKEYALFY RLDIVPLNEN SSEYRLINCN T STITQICP ...String: NLWVTVYYGV PVWKEAKTTL FCASDAKAYE KEVHNVWATH ACVPTDPNPQ EMVLENVTEN FNMWKNDMVD QMHEDIISLW DQSLKPCVK LTPLCVTLNC TNVNVTNTNN NNMKEEMKNC SFNTTTEIRD KKQKEYALFY RLDIVPLNEN SSEYRLINCN T STITQICP KVSFDPIPIH YCAPAGYAIL KCNNKTFNGT GPCNNVSTVQ CTHGIKPVVS TQLLLNGSLA EEEIIIRSEN LT DNAKTII VHLNESVEIN CTRPNNMTRK SIRIGPGQTF YALGDIIGDI RQPHCNISEA KWNKTLQRVK KKLKEHFPNK TIK FAPSSG GDLEITTHSF NCRGEFFYCN TSKLFNSTYN NTTSNSTITL PCRIKQIINM WQEVGRAMYA PPIAGNITCK SNIT GLLLT RDGGNNNNNT ETFRPGGGDM RDNWRSELYK YKVVEIKPLG IAPTKCKRRV VERRRRRR |
-Macromolecule #2: TRNM-f*01 heavy chain
| Macromolecule | Name: TRNM-f*01 heavy chain / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.923301 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLQESGPG VVKPSETLSL TCAVSGDSIK SAYQYWNWIR QPRGKGPEWI GGVYSSSDST AYNPSLESRV SISRDTSNNR FSLNLRSVT ATDTATYFCA RSVRDSRGWG RYFLDTWGQG LLVTVSS |
-Macromolecule #3: TRNM-f*01 light chain
| Macromolecule | Name: TRNM-f*01 light chain / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 11.464811 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DIQMTQSPSS LSASVGDTVT ITCRASQSIS TWLAWYQQKP GKAPKVLIYS ASILQSGVPS RFRGSGSGSD FTLTIGSLQI EDFATYFCQ QYTGSPFTFG GGTKVEIK |
-Macromolecule #4: Envelope glycoprotein gp41
| Macromolecule | Name: Envelope glycoprotein gp41 / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 17.174447 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AVGLGAVFLG FLGAAGSTMG AASNTLTVQA RQLLSGIVQQ QSNLLRAPEA QQHMLQLGVW GFKQLQARVL AIERYLEVQQ LLGIWGCSG KLICCTAVPW NSSWSNKSQE DIWDNMTWMQ WDREIGNYTD TIYRLLEESQ FQQEINEKDL LALD |
-Macromolecule #7: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 7 / Number of copies: 42 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 / Details: PBS |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 58.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)