+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Truncated Braf/Mek/14-3-3 complex | ||||||||||||
 Map data Map data | (del1-155)Braf/Mek/14-3-3 complex | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | kinase complex / TRANSFERASE | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.1 Å | ||||||||||||
 Authors Authors | Eck MJ / Jeon H / Park E / Rawson S | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Cryo-EM structure of a RAS/RAF recruitment complex. Authors: Eunyoung Park / Shaun Rawson / Anna Schmoker / Byeong-Won Kim / Sehee Oh / Kangkang Song / Hyesung Jeon / Michael J Eck /   Abstract: RAF-family kinases are activated by recruitment to the plasma membrane by GTP-bound RAS, whereupon they initiate signaling through the MAP kinase cascade. Prior structural studies of KRAS with RAF ...RAF-family kinases are activated by recruitment to the plasma membrane by GTP-bound RAS, whereupon they initiate signaling through the MAP kinase cascade. Prior structural studies of KRAS with RAF have focused on the isolated RAS-binding and cysteine-rich domains of RAF (RBD and CRD, respectively), which interact directly with RAS. Here we describe cryo-EM structures of a KRAS bound to intact BRAF in an autoinhibited state with MEK1 and a 14-3-3 dimer. Analysis of this KRAS/BRAF/MEK1/14-3-3 complex reveals KRAS bound to the RAS-binding domain of BRAF, captured in two orientations. Core autoinhibitory interactions in the complex are unperturbed by binding of KRAS and in vitro activation studies confirm that KRAS binding is insufficient to activate BRAF, absent membrane recruitment. These structures illustrate the separability of binding and activation of BRAF by RAS and suggest stabilization of this pre-activation intermediate as an alternative therapeutic strategy to blocking binding of KRAS. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40253.map.gz emd_40253.map.gz | 28.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40253-v30.xml emd-40253-v30.xml emd-40253.xml emd-40253.xml | 18.1 KB 18.1 KB | Display Display |  EMDB header EMDB header |
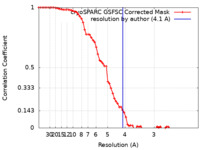
| FSC (resolution estimation) |  emd_40253_fsc.xml emd_40253_fsc.xml | 6.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_40253.png emd_40253.png | 50 KB | ||
| Masks |  emd_40253_msk_1.map emd_40253_msk_1.map | 30.5 MB |  Mask map Mask map | |
| Others |  emd_40253_additional_1.map.gz emd_40253_additional_1.map.gz emd_40253_half_map_1.map.gz emd_40253_half_map_1.map.gz emd_40253_half_map_2.map.gz emd_40253_half_map_2.map.gz | 21.1 MB 28.3 MB 28.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40253 http://ftp.pdbj.org/pub/emdb/structures/EMD-40253 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40253 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40253 | HTTPS FTP |
-Validation report
| Summary document |  emd_40253_validation.pdf.gz emd_40253_validation.pdf.gz | 802.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_40253_full_validation.pdf.gz emd_40253_full_validation.pdf.gz | 802.1 KB | Display | |
| Data in XML |  emd_40253_validation.xml.gz emd_40253_validation.xml.gz | 14.2 KB | Display | |
| Data in CIF |  emd_40253_validation.cif.gz emd_40253_validation.cif.gz | 18 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40253 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40253 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40253 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40253 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_40253.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40253.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | (del1-155)Braf/Mek/14-3-3 complex | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_40253_msk_1.map emd_40253_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: (del1-155)Braf/Mek/14-3-3 complex
| File | emd_40253_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | (del1-155)Braf/Mek/14-3-3 complex | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: (del1-155)Braf/Mek/14-3-3 complex
| File | emd_40253_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | (del1-155)Braf/Mek/14-3-3 complex | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: (del1-155)Braf/Mek/14-3-3 complex
| File | emd_40253_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | (del1-155)Braf/Mek/14-3-3 complex | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Truncated Braf/Mek/14-3-3 complex
| Entire | Name: Truncated Braf/Mek/14-3-3 complex |
|---|---|
| Components |
|
-Supramolecule #1: Truncated Braf/Mek/14-3-3 complex
| Supramolecule | Name: Truncated Braf/Mek/14-3-3 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 / Details: (del1-155)Braf/Mek/14-3-3 complex |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 185 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Component - Concentration: 150.0 mM / Component - Formula: NaCl / Component - Name: sodium chloride Details: 50 mM Tris pH 7.4, 150 mM NaCl, 2 mM MgCl2, 0.5 mM TCEP, 50 uM ATP-gammaS |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV |
| Details | monodisperse |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Phase plate: VOLTA PHASE PLATE |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number grids imaged: 1 / Number real images: 3252 / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.2 µm / Nominal defocus min: 0.2 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER |
|---|
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)