+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the human endogenous PCNA-FEN1 complex - State A | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Flap endonuclease 1 / endogenous DNA / PCNA / DNA BINDING PROTEIN/DNA / DNA BINDING PROTEIN-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationflap endonuclease activity / positive regulation of sister chromatid cohesion / double-stranded DNA exodeoxyribonuclease activity / telomere maintenance via semi-conservative replication / dinucleotide insertion or deletion binding / PCNA-p21 complex / mitotic telomere maintenance via semi-conservative replication / 5'-flap endonuclease activity / purine-specific mismatch base pair DNA N-glycosylase activity / nuclear lamina ...flap endonuclease activity / positive regulation of sister chromatid cohesion / double-stranded DNA exodeoxyribonuclease activity / telomere maintenance via semi-conservative replication / dinucleotide insertion or deletion binding / PCNA-p21 complex / mitotic telomere maintenance via semi-conservative replication / 5'-flap endonuclease activity / purine-specific mismatch base pair DNA N-glycosylase activity / nuclear lamina / Polymerase switching / Processive synthesis on the lagging strand / MutLalpha complex binding / PCNA complex / DNA replication, removal of RNA primer / Telomere C-strand (Lagging Strand) Synthesis / Removal of the Flap Intermediate / UV protection / HDR through MMEJ (alt-NHEJ) / Mismatch repair (MMR) directed by MSH2:MSH3 (MutSbeta) / Mismatch repair (MMR) directed by MSH2:MSH6 (MutSalpha) / Transcription of E2F targets under negative control by DREAM complex / Polymerase switching on the C-strand of the telomere / replisome / Processive synthesis on the C-strand of the telomere / 5'-3' exonuclease activity / response to L-glutamate / Removal of the Flap Intermediate from the C-strand / exonuclease activity / response to dexamethasone / DNA polymerase processivity factor activity / histone acetyltransferase binding / Early Phase of HIV Life Cycle / leading strand elongation / G1/S-Specific Transcription / nuclear replication fork / replication fork processing / SUMOylation of DNA replication proteins / POLB-Dependent Long Patch Base Excision Repair / PCNA-Dependent Long Patch Base Excision Repair / response to cadmium ion / estrous cycle / mismatch repair / cyclin-dependent protein kinase holoenzyme complex / translesion synthesis / base-excision repair, gap-filling / DNA polymerase binding / epithelial cell differentiation / liver regeneration / TP53 Regulates Transcription of Genes Involved in G2 Cell Cycle Arrest / positive regulation of DNA replication / Translesion synthesis by REV1 / nuclear estrogen receptor binding / positive regulation of DNA repair / Translesion synthesis by POLK / replication fork / Translesion synthesis by POLI / Gap-filling DNA repair synthesis and ligation in GG-NER / male germ cell nucleus / Termination of translesion DNA synthesis / Translesion Synthesis by POLH / Recognition of DNA damage by PCNA-containing replication complex / receptor tyrosine kinase binding / double-strand break repair via homologous recombination / HDR through Homologous Recombination (HRR) / cellular response to xenobiotic stimulus / Dual Incision in GG-NER / RNA-DNA hybrid ribonuclease activity / cellular response to hydrogen peroxide / memory / Dual incision in TC-NER / Gap-filling DNA repair synthesis and ligation in TC-NER / cellular response to UV / response to estradiol / double-strand break repair / manganese ion binding / E3 ubiquitin ligases ubiquitinate target proteins / heart development / chromatin organization / double-stranded DNA binding / endonuclease activity / damaged DNA binding / Hydrolases; Acting on ester bonds / DNA replication / chromosome, telomeric region / nuclear body / DNA repair / chromatin binding / centrosome / chromatin / protein-containing complex binding / nucleolus / magnesium ion binding / enzyme binding / negative regulation of transcription by RNA polymerase II / protein-containing complex / mitochondrion / DNA binding / extracellular exosome / nucleoplasm Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
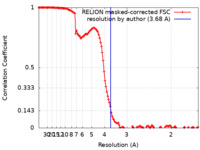
| Method | single particle reconstruction / cryo EM / Resolution: 3.68 Å | |||||||||
 Authors Authors | Tian Y / Gao N | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: EMBO J / Year: 2025 Journal: EMBO J / Year: 2025Title: Structural insight into Okazaki fragment maturation mediated by PCNA-bound FEN1 and RNaseH2. Authors: Yuhui Tian / Ningning Li / Qing Li / Ning Gao /  Abstract: PCNA is a master coordinator of many DNA-metabolic events. During DNA replication, the maturation of Okazaki fragments involves at least four DNA enzymes, all of which contain PCNA-interacting motifs. ...PCNA is a master coordinator of many DNA-metabolic events. During DNA replication, the maturation of Okazaki fragments involves at least four DNA enzymes, all of which contain PCNA-interacting motifs. However, the temporal relationships and functional modulations between these PCNA-binding proteins are unclear. Here, we developed a strategy to purify endogenous PCNA-containing complexes from native chromatin, and characterized their structures using cryo-EM. Two structurally resolved classes (PCNA-FEN1 and PCNA-FEN1-RNaseH2 complexes) have captured a series of 3D snapshots for the primer-removal steps of Okazaki fragment maturation. These structures show that product release from FEN1 is a rate-liming step. Furthermore, both FEN1 and RNaseH2 undergo continuous conformational changes on PCNA that result in constant fluctuations in the bending angle of substrate DNA at the nick site, implying that these enzymes could regulate each other through conformational modulation of the bound DNA. The structures of the PCNA-FEN1-RNaseH2 complex confirm the toolbelt function of PCNA and suggests a potential unrecognized role of RNaseH2, as a dsDNA binding protein, in promoting the 5'-flap cleaving activity of FEN1. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_39342.map.gz emd_39342.map.gz | 10 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-39342-v30.xml emd-39342-v30.xml emd-39342.xml emd-39342.xml | 19.8 KB 19.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_39342_fsc.xml emd_39342_fsc.xml | 12.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_39342.png emd_39342.png | 99.6 KB | ||
| Filedesc metadata |  emd-39342.cif.gz emd-39342.cif.gz | 6.4 KB | ||
| Others |  emd_39342_half_map_1.map.gz emd_39342_half_map_1.map.gz emd_39342_half_map_2.map.gz emd_39342_half_map_2.map.gz | 141.3 MB 140.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-39342 http://ftp.pdbj.org/pub/emdb/structures/EMD-39342 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39342 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39342 | HTTPS FTP |
-Related structure data
| Related structure data |  8yjhMC  8yjlC  8yjqC  8yjrC  8yjsC  8yjuC  8yjvC  8yjwC  8yjzC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_39342.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_39342.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_39342_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_39342_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : endogenous state A PCNA-DNA-FEN1 complex
| Entire | Name: endogenous state A PCNA-DNA-FEN1 complex |
|---|---|
| Components |
|
-Supramolecule #1: endogenous state A PCNA-DNA-FEN1 complex
| Supramolecule | Name: endogenous state A PCNA-DNA-FEN1 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Proliferating cell nuclear antigen
| Macromolecule | Name: Proliferating cell nuclear antigen / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 28.795752 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFEARLVQGS ILKKVLEALK DLINEACWDI SSSGVNLQSM DSSHVSLVQL TLRSEGFDTY RCDRNLAMGV NLTSMSKILK CAGNEDIIT LRAEDNADTL ALVFEAPNQE KVSDYEMKLM DLDVEQLGIP EQEYSCVVKM PSGEFARICR DLSHIGDAVV I SCAKDGVK ...String: MFEARLVQGS ILKKVLEALK DLINEACWDI SSSGVNLQSM DSSHVSLVQL TLRSEGFDTY RCDRNLAMGV NLTSMSKILK CAGNEDIIT LRAEDNADTL ALVFEAPNQE KVSDYEMKLM DLDVEQLGIP EQEYSCVVKM PSGEFARICR DLSHIGDAVV I SCAKDGVK FSASGELGNG NIKLSQTSNV DKEEEAVTIE MNEPVQLTFA LRYLNFFTKA TPLSSTVTLS MSADVPLVVE YK IADMGHL KYYLAPKIED EEGS UniProtKB: DNA sliding clamp PCNA |
-Macromolecule #2: Flap endonuclease 1
| Macromolecule | Name: Flap endonuclease 1 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: Hydrolases; Acting on ester bonds |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 42.661051 KDa |
| Sequence | String: MGIQGLAKLI ADVAPSAIRE NDIKSYFGRK VAIDASMSIY QFLIAVRQGG DVLQNEEGET TSHLMGMFYR TIRMMENGIK PVYVFDGKP PQLKSGELAK RSERRAEAEK QLQQAQAAGA EQEVEKFTKR LVKVTKQHND ECKHLLSLMG IPYLDAPSEA E ASCAALVK ...String: MGIQGLAKLI ADVAPSAIRE NDIKSYFGRK VAIDASMSIY QFLIAVRQGG DVLQNEEGET TSHLMGMFYR TIRMMENGIK PVYVFDGKP PQLKSGELAK RSERRAEAEK QLQQAQAAGA EQEVEKFTKR LVKVTKQHND ECKHLLSLMG IPYLDAPSEA E ASCAALVK AGKVYAAATE DMDCLTFGSP VLMRHLTASE AKKLPIQEFH LSRILQELGL NQEQFVDLCI LLGSDYCESI RG IGPKRAV DLIQKHKSIE EIVRRLDPNK YPVPENWLHK EAHQLFLEPE VLDPESVELK WSEPNEEELI KFMCGEKQFS EER IRSGVK RLSKSRQGST QGRLDDFFKV TGSLSSAKRK EPEPKGSTKK KAKTGAAGKF KRGK UniProtKB: Flap endonuclease 1 |
-Macromolecule #3: upstream DNA chain J
| Macromolecule | Name: upstream DNA chain J / type: dna / ID: 3 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 4.590046 KDa |
| Sequence | String: (DT)(DA)(DA)(DA)(DA)(DA)(DT)(DT)(DT)(DA) (DA)(DA)(DT)(DT)(DT) |
-Macromolecule #4: parent strand chain E
| Macromolecule | Name: parent strand chain E / type: dna / ID: 4 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.668031 KDa |
| Sequence | String: (DT)(DT)(DT)(DT)(DT)(DA)(DA)(DA)(DA)(DA) (DA)(DA)(DA)(DT)(DT)(DT)(DA)(DA)(DA)(DT) (DT)(DT)(DT)(DT)(DA) |
-Macromolecule #5: downstream DNA chain F
| Macromolecule | Name: downstream DNA chain F / type: dna / ID: 5 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 3.042041 KDa |
| Sequence | String: (DT)(DT)(DT)(DT)(DT)(DA)(DA)(DA)(DA)(DA) |
-Macromolecule #6: 5 prime DNA
| Macromolecule | Name: 5 prime DNA / type: dna / ID: 6 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 867.621 Da |
| Sequence | String: (DT)(DT)(DT) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 2.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)