[English] 日本語
 Yorodumi
Yorodumi- EMDB-39288: Cryo-EM structure of CTR-bound type 7 CRISPR-Cas complex at post-... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of CTR-bound type 7 CRISPR-Cas complex at post-state 2 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | protein structure / ANTIVIRAL PROTEIN | |||||||||
| Biological species | metagenome (others) | |||||||||
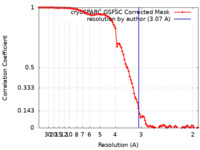
| Method | single particle reconstruction / cryo EM / Resolution: 3.07 Å | |||||||||
 Authors Authors | Zhang H / Deng Z / Li X | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Cryo-EM structure of CTR-bound type 7 CRISPR-Cas complex at post-state 2 Authors: Zhang H / Deng Z / Li X | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_39288.map.gz emd_39288.map.gz | 97.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-39288-v30.xml emd-39288-v30.xml emd-39288.xml emd-39288.xml | 16.5 KB 16.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_39288_fsc.xml emd_39288_fsc.xml | 9.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_39288.png emd_39288.png | 82 KB | ||
| Filedesc metadata |  emd-39288.cif.gz emd-39288.cif.gz | 5.9 KB | ||
| Others |  emd_39288_half_map_1.map.gz emd_39288_half_map_1.map.gz emd_39288_half_map_2.map.gz emd_39288_half_map_2.map.gz | 95.5 MB 95.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-39288 http://ftp.pdbj.org/pub/emdb/structures/EMD-39288 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39288 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39288 | HTTPS FTP |
-Related structure data
| Related structure data |  8yhe M: atomic model generated by this map |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_39288.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_39288.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.95 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_39288_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_39288_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : protein structure
| Entire | Name: protein structure |
|---|---|
| Components |
|
-Supramolecule #1: protein structure
| Supramolecule | Name: protein structure / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism: metagenome (others) |
-Macromolecule #1: protein structure
| Macromolecule | Name: protein structure / type: protein_or_peptide / ID: 1 / Number of copies: 7 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: metagenome (others) |
| Molecular weight | Theoretical: 22.255604 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAKTMKKIYV TMKTLSPLYT GEVRREDKEA AQKRVNFPVR KTATNKVLIP FKGALRSALE IMLKAKGENV CDTGESRARP CGRCVTCSL FGSMGRAGRA SVDFLISNDT KEQIVRESTH LRIERQTKSA SDTFKGEEVI EGATFTATIT ISNPQEKDLS L IQSALKFI ...String: MAKTMKKIYV TMKTLSPLYT GEVRREDKEA AQKRVNFPVR KTATNKVLIP FKGALRSALE IMLKAKGENV CDTGESRARP CGRCVTCSL FGSMGRAGRA SVDFLISNDT KEQIVRESTH LRIERQTKSA SDTFKGEEVI EGATFTATIT ISNPQEKDLS L IQSALKFI EENGIGGWLN KGYGRVSFEV KSEDVATDRF LK |
-Macromolecule #2: protein structure
| Macromolecule | Name: protein structure / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: metagenome (others) |
| Molecular weight | Theoretical: 27.139533 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKEIKGILES ITGFSIPLDN GEYALYPAGR HLRGAIGYIA FNLDLPISSK FLDFDFDDII FRDLLPISKC GKIFYPEKNS NSLKCPSCN EIYGSSVLRN IMARGLSYKE VIEGKKYRLS IIVKDEKYLN EMEAIIRYIL SYGIYLGNKV SKGYGKFKIK E YSIVDILP ...String: MKEIKGILES ITGFSIPLDN GEYALYPAGR HLRGAIGYIA FNLDLPISSK FLDFDFDDII FRDLLPISKC GKIFYPEKNS NSLKCPSCN EIYGSSVLRN IMARGLSYKE VIEGKKYRLS IIVKDEKYLN EMEAIIRYIL SYGIYLGNKV SKGYGKFKIK E YSIVDILP VKDSEVLLLS DAIIDNGEKD IVFSKKEISS SKFEIIRKRG KAKGDIIRDN NHNGFYIGKY GGLGFGEIIS LK |
-Macromolecule #3: protein structure
| Macromolecule | Name: protein structure / type: protein_or_peptide / ID: 3 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: metagenome (others) |
| Molecular weight | Theoretical: 70.465875 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MIKFIGGASK VTGSAFLLET GAKILIDCGI EQEKGIEKDN NEIIEKKINE IGKADICILT HAHLDHSGLV PLLVKKRKVN KIISTPATK ELCRLLFNDF QRIQEENNDI PLYSYDDIES SFEIWDEIDD RNTIELFDTK ITFYNNSHII GSVSVFIETH N GNYLFSGD ...String: MIKFIGGASK VTGSAFLLET GAKILIDCGI EQEKGIEKDN NEIIEKKINE IGKADICILT HAHLDHSGLV PLLVKKRKVN KIISTPATK ELCRLLFNDF QRIQEENNDI PLYSYDDIES SFEIWDEIDD RNTIELFDTK ITFYNNSHII GSVSVFIETH N GNYLFSGD IGSKLQQLMD YPPDMPDGNV DYLILESTYG NKSHDSSDRD RLLEIAKTTC ENGGKVLIPS FAIGRLQEVL YT FSNYNFN FPVYIDSPMG SKVTNLIKEY NIYLKKKLRR LSITDDLFNN KYIAINTSNQ SKELSNSKEP AVIISASGML EGG RILNHL EQIKNDENST LIFVGYQAQN TRGRKILDGE EKVRCRIEKL NSFSAHADQD ELIDYIERLK YTPYKVFLVH GEKE QREIL AKRIISKKIR VELPENYSQG KEILIEKKVV LNINTDNMCN FASYRLMPFS GFIVEKDDRI EINDKNWFDM IWNEE YNKM RSQIVAEDFS TDQNEDSMAL PDMSHDKIIE NIEYLFNIKI LSKNRIKEFW EEFCKGQKAA IKYITQVHRK NPNTGR RNW NPPEGDFTDN EIEKLYETAY NTLLSLIKYD KNKVYNILIN FNPKL |
-Macromolecule #4: RNA (46-MER)
| Macromolecule | Name: RNA (46-MER) / type: rna / ID: 4 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism: metagenome (others) |
| Molecular weight | Theoretical: 15.300997 KDa |
| Sequence | String: GUUAAAACUC UUCUCAUGCU GGAUUCGAAA UUAGGUGCGC UUCGCGUU |
-Macromolecule #5: RNA (29-MER)
| Macromolecule | Name: RNA (29-MER) / type: rna / ID: 5 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism: metagenome (others) |
| Molecular weight | Theoretical: 16.713082 KDa |
| Sequence | String: GAACACCCAA UAGCGAAGCG CACCUAAUUU CGAAUCCAGC AUGAGAAGCU AA |
-Macromolecule #6: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 6 / Number of copies: 9 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | JEOL CRYO ARM 300 |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm |
 Movie
Movie Controller
Controller


 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)