+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the 123-316 scDb/PT-RBD complex | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Cryo-EM / SARS-CoV2 / antibody / RBD / IMMUNE SYSTEM | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell ...symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell / membrane fusion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / Attachment and Entry / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / endocytosis involved in viral entry into host cell / receptor ligand activity / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / membrane / identical protein binding / plasma membrane Similarity search - Function | |||||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.86 Å | |||||||||||||||
 Authors Authors | Jia GW / Tong Z / Tong JY / Su ZM | |||||||||||||||
| Funding support |  China, 4 items China, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2024 Journal: Cell Rep / Year: 2024Title: Deciphering a reliable synergistic bispecific strategy of rescuing antibodies for SARS-CoV-2 escape variants, including BA.2.86, EG.5.1, and JN.1. Authors: Zhou Tong / Jianyu Tong / Wenwen Lei / Yufeng Xie / Yingzi Cui / Guowen Jia / Shihua Li / Zezhong Zhang / Zhimin Cheng / Xiao Xing / Haiyun Ma / Lan Deng / Rong Zhang / Xin Zhao / Kefang Liu ...Authors: Zhou Tong / Jianyu Tong / Wenwen Lei / Yufeng Xie / Yingzi Cui / Guowen Jia / Shihua Li / Zezhong Zhang / Zhimin Cheng / Xiao Xing / Haiyun Ma / Lan Deng / Rong Zhang / Xin Zhao / Kefang Liu / Qihui Wang / Jianxun Qi / Haomin Huang / Rui Song / Zhaoming Su / Guizhen Wu / Jing Lou / George Fu Gao /  Abstract: The game between therapeutic monoclonal antibodies (mAbs) and continuously emerging severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) variants has favored the virus, as most therapeutic ...The game between therapeutic monoclonal antibodies (mAbs) and continuously emerging severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) variants has favored the virus, as most therapeutic mAbs have been evaded. Addressing this challenge, we systematically explored a reproducible bispecific antibody (bsAb)-dependent synergistic effect in this study. It could effectively restore the neutralizing activity of the bsAb when any of its single mAbs is escaped by variants. This synergy is primarily attributed to the binding angle of receptor-binding domain (RBD)-5, facilitating inter-spike cross-linking and promoting cryptic epitope exposure that classical antibody cocktails cannot achieve. Furthermore, RBD-5 with RBD-2, RBD-6, and RBD-7, alongside RBD-8, also exhibit significantly enhanced effects. This study not only shifts the paradigm in understanding antibody interactions but paves the way for developing more effective therapeutic antibodies against rapidly mutating SARS-CoV-2, with Dia-19 already showing promise against emerging variants like BA.2.86, EG.5.1, and JN.1. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_38823.map.gz emd_38823.map.gz | 85.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-38823-v30.xml emd-38823-v30.xml emd-38823.xml emd-38823.xml | 18.7 KB 18.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_38823_fsc.xml emd_38823_fsc.xml | 9.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_38823.png emd_38823.png | 61.8 KB | ||
| Filedesc metadata |  emd-38823.cif.gz emd-38823.cif.gz | 6.4 KB | ||
| Others |  emd_38823_half_map_1.map.gz emd_38823_half_map_1.map.gz emd_38823_half_map_2.map.gz emd_38823_half_map_2.map.gz | 84.7 MB 84.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-38823 http://ftp.pdbj.org/pub/emdb/structures/EMD-38823 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38823 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-38823 | HTTPS FTP |
-Related structure data
| Related structure data |  8y0yMC  8xseC  8xsfC  8xsiC  8xsjC  8xslC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_38823.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_38823.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||
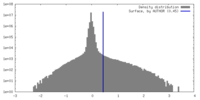
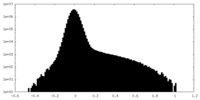
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_38823_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
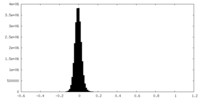
| Density Histograms |
-Half map: #1
| File | emd_38823_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
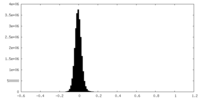
| Density Histograms |
- Sample components
Sample components
-Entire : The 123-316 scDb/PT-RBD complex
| Entire | Name: The 123-316 scDb/PT-RBD complex |
|---|---|
| Components |
|
-Supramolecule #1: The 123-316 scDb/PT-RBD complex
| Supramolecule | Name: The 123-316 scDb/PT-RBD complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: 123-316 scDb
| Macromolecule | Name: 123-316 scDb / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 56.931738 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: METDTLLLWV LLLWVPGSTG DQVQLVESGG GVVQPGRSLR LSCAASGFTF SNYGMHWVRQ APGKGLEWVA VISHDASNKY YADSVKGRF TISRDNSKNT QYLQMNSLRA EDTAVYYCAK GGGYSYVVDM GPYWGQGTLV TVSSASRADS AAAGGPGSDI Q MTQSPSSV ...String: METDTLLLWV LLLWVPGSTG DQVQLVESGG GVVQPGRSLR LSCAASGFTF SNYGMHWVRQ APGKGLEWVA VISHDASNKY YADSVKGRF TISRDNSKNT QYLQMNSLRA EDTAVYYCAK GGGYSYVVDM GPYWGQGTLV TVSSASRADS AAAGGPGSDI Q MTQSPSSV SASVGDSVTI TCRASQGISR WLAWYQQRPG KAPKLLIYAA GNLETGVPSR FSGSGSGTDF TLTISDLQAE DF ATYYCQQ ADSFPLTFGG GTKVDIKGRA DAAAAGGGGS GGGGSGGGGS GGGGSQVQLV ESGGGLVQPG GSLRLSCAAS GFT FSSYAM SWVRQAPGKG LEWVSAISGS GGSTYYADSV KGRFTISRDN SKNTLYLQMN SLRAEDTAVY YCAKDHLITM VQPE YFHHW GQGTLVTVSS ASRADAAAAG GPGDIQMTQS PSSLSASVGD RVTITCRASQ SISNYLNWYQ QKPGKAPKLL IYDAS NLET GVPSRFSGSG SGADFTFTIG SLQPEDSATY YCQQYDNLPL TFGGGTKVEI KGHHHHHH |
-Macromolecule #2: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 23.513389 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: RVQPTESIVR FPNITNLCPF GEVFNATRFA SVYAWNRKRI SNCVADYSVL YNSASFSTFK CYGVSPTKLN DLCFTNVYAD SFVIRGDEV RQIAPGQTGK IADYNYKLPD DFTGCVIAWN SNNLDSKVGG NYNYLYRLFR KSNLKPFERD ISTEIYQAGS T PCNGVEGF ...String: RVQPTESIVR FPNITNLCPF GEVFNATRFA SVYAWNRKRI SNCVADYSVL YNSASFSTFK CYGVSPTKLN DLCFTNVYAD SFVIRGDEV RQIAPGQTGK IADYNYKLPD DFTGCVIAWN SNNLDSKVGG NYNYLYRLFR KSNLKPFERD ISTEIYQAGS T PCNGVEGF NCYFPLQSYG FQPTNGVGYQ PYRVVVLSFE LLHAPATVCG P UniProtKB: Spike glycoprotein |
-Macromolecule #3: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 3 / Number of copies: 1 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 53.1 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.1 µm / Nominal defocus min: 0.3 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)