+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | SID1 transmembrane family member 2 | |||||||||
 マップデータ マップデータ | ||||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | VIRAL PROTEIN-IMMUNE SYSTEM COMPLEX / TRANSPORT PROTEIN | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報nucleic acid transmembrane transporter activity / AP-1 adaptor complex binding / RNA transmembrane transporter activity / AP-2 adaptor complex binding / RNA transport / type B pancreatic cell development / regulation of insulin secretion involved in cellular response to glucose stimulus / type B pancreatic cell proliferation / RNA catabolic process / response to glucose ...nucleic acid transmembrane transporter activity / AP-1 adaptor complex binding / RNA transmembrane transporter activity / AP-2 adaptor complex binding / RNA transport / type B pancreatic cell development / regulation of insulin secretion involved in cellular response to glucose stimulus / type B pancreatic cell proliferation / RNA catabolic process / response to glucose / cell morphogenesis / double-stranded RNA binding / glucose homeostasis / lysosome / lysosomal membrane / DNA binding / plasma membrane 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) Homo sapiens (ヒト) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.3 Å | |||||||||
 データ登録者 データ登録者 | Guo H / Qi C / Lu Y / Yang H / Zhu Y / Sun F / Ji X | |||||||||
| 資金援助 |  中国, 1件 中国, 1件
| |||||||||
 引用 引用 |  ジャーナル: Cell Res / 年: 2024 ジャーナル: Cell Res / 年: 2024タイトル: Cryo-EM structures of human SID-1 transmembrane family proteins and implications for their low-pH-dependent RNA transport activity. 著者: Le Zheng / Tingting Yang / Hangtian Guo / Chen Qi / Yuchi Lu / Haonan Xiao / Yan Gao / Yue Liu / Yixuan Yang / Mengru Zhou / Henry C Nguyen / Yun Zhu / Fei Sun / Chen-Yu Zhang / Xiaoyun Ji /  | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_36784.map.gz emd_36784.map.gz | 117.9 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-36784-v30.xml emd-36784-v30.xml emd-36784.xml emd-36784.xml | 14.1 KB 14.1 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_36784.png emd_36784.png | 39.7 KB | ||
| Filedesc metadata |  emd-36784.cif.gz emd-36784.cif.gz | 5.5 KB | ||
| その他 |  emd_36784_half_map_1.map.gz emd_36784_half_map_1.map.gz emd_36784_half_map_2.map.gz emd_36784_half_map_2.map.gz | 115.7 MB 115.7 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36784 http://ftp.pdbj.org/pub/emdb/structures/EMD-36784 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36784 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36784 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_36784.map.gz / 形式: CCP4 / 大きさ: 125 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_36784.map.gz / 形式: CCP4 / 大きさ: 125 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 0.82 Å | ||||||||||||||||||||||||||||||||||||
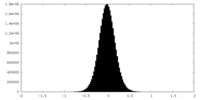
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-ハーフマップ: #2
| ファイル | emd_36784_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
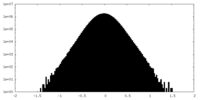
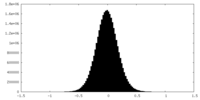
| 密度ヒストグラム |
-ハーフマップ: #1
| ファイル | emd_36784_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
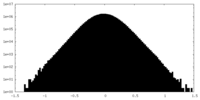
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : human SID1 transmembrane family member 2 (SIDT2) homodimer, TMD f...
| 全体 | 名称: human SID1 transmembrane family member 2 (SIDT2) homodimer, TMD focused refinement |
|---|---|
| 要素 |
|
-超分子 #1: human SID1 transmembrane family member 2 (SIDT2) homodimer, TMD f...
| 超分子 | 名称: human SID1 transmembrane family member 2 (SIDT2) homodimer, TMD focused refinement タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: all |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
-分子 #1: SID1 transmembrane family member 2
| 分子 | 名称: SID1 transmembrane family member 2 / タイプ: protein_or_peptide / ID: 1 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 60.065715 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: VTSEAYVSGM LFCLGIFLSF YLLTVLLACW ENWRQKKKTL LVAIDRACPE SGHPRVLADS FPGSSPYEGY NYGSFENVDT LTDIDSDKN VIRTKQYLYV ADLARKDKRV LRKKYQIYFW NIATIAVFYA LPVVQLVITY QTVVNVTGNQ DICYYNFLCA H PLGNLSAF ...文字列: VTSEAYVSGM LFCLGIFLSF YLLTVLLACW ENWRQKKKTL LVAIDRACPE SGHPRVLADS FPGSSPYEGY NYGSFENVDT LTDIDSDKN VIRTKQYLYV ADLARKDKRV LRKKYQIYFW NIATIAVFYA LPVVQLVITY QTVVNVTGNQ DICYYNFLCA H PLGNLSAF NNILSNLGYI LLGLLFLLII LQREINHNRA LLRNDLCALE CGIPKHFGLF YAMGTALMME GLLSACYHVC PN YTNFQFD TSFMYMIAGL CMLKLYQKRH PDINASAYSA YACLAIVIFF SVLGVVFGKG NTAFWIVFSI IHIIATLLLS TQL YYMGRW KLDSGIFRRI LHVLYTDCIR QCSGPLYVDR MVLLVMGNVI NWSLAAYGLI MRPNDFASYL LAIGICNLLL YFAF YIIMK LRSGERIKLI PLLCIVCTSV VWGFALFFFF QGLSTWQKTP AESREHNRDC ILLDFFDDHD IWHFLSSIAM FGSFL VLLT LDDDLDTVQR DKIYVFDYKD HDGDYKDHDI DYKDDDDK UniProtKB: SID1 transmembrane family member 2 |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.5 |
|---|---|
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 281 K / 装置: FEI VITROBOT MARK IV |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K3 (6k x 4k) / 平均電子線量: 60.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: SPOT SCAN / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 2.0 µm / 最小 デフォーカス(公称値): 1.2 µm |
- 画像解析
画像解析
| 初期モデル | モデルのタイプ: NONE |
|---|---|
| 最終 再構成 | 解像度のタイプ: BY AUTHOR / 解像度: 3.3 Å / 解像度の算出法: FSC 0.143 CUT-OFF / 使用した粒子像数: 67365 |
| 初期 角度割当 | タイプ: COMMON LINE |
| 最終 角度割当 | タイプ: COMMON LINE |
 ムービー
ムービー コントローラー
コントローラー














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)