+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | SID1 transmembrane family member 2 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | VIRAL PROTEIN-IMMUNE SYSTEM COMPLEX / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationnucleic acid transmembrane transporter activity / AP-1 adaptor complex binding / RNA transmembrane transporter activity / AP-2 adaptor complex binding / RNA transport / type B pancreatic cell development / regulation of insulin secretion involved in cellular response to glucose stimulus / type B pancreatic cell proliferation / RNA catabolic process / response to glucose ...nucleic acid transmembrane transporter activity / AP-1 adaptor complex binding / RNA transmembrane transporter activity / AP-2 adaptor complex binding / RNA transport / type B pancreatic cell development / regulation of insulin secretion involved in cellular response to glucose stimulus / type B pancreatic cell proliferation / RNA catabolic process / response to glucose / cell morphogenesis / double-stranded RNA binding / glucose homeostasis / lysosome / lysosomal membrane / DNA binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Guo H / Qi C / Lu Y / Yang H / Zhu Y / Sun F / Ji X | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Res / Year: 2024 Journal: Cell Res / Year: 2024Title: Cryo-EM structures of human SID-1 transmembrane family proteins and implications for their low-pH-dependent RNA transport activity. Authors: Le Zheng / Tingting Yang / Hangtian Guo / Chen Qi / Yuchi Lu / Haonan Xiao / Yan Gao / Yue Liu / Yixuan Yang / Mengru Zhou / Henry C Nguyen / Yun Zhu / Fei Sun / Chen-Yu Zhang / Xiaoyun Ji /  | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36783.map.gz emd_36783.map.gz | 117.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36783-v30.xml emd-36783-v30.xml emd-36783.xml emd-36783.xml | 14.3 KB 14.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_36783.png emd_36783.png | 42.5 KB | ||
| Filedesc metadata |  emd-36783.cif.gz emd-36783.cif.gz | 5.5 KB | ||
| Others |  emd_36783_half_map_1.map.gz emd_36783_half_map_1.map.gz emd_36783_half_map_2.map.gz emd_36783_half_map_2.map.gz | 115.7 MB 115.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36783 http://ftp.pdbj.org/pub/emdb/structures/EMD-36783 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36783 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36783 | HTTPS FTP |
-Validation report
| Summary document |  emd_36783_validation.pdf.gz emd_36783_validation.pdf.gz | 914.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_36783_full_validation.pdf.gz emd_36783_full_validation.pdf.gz | 914.5 KB | Display | |
| Data in XML |  emd_36783_validation.xml.gz emd_36783_validation.xml.gz | 13.9 KB | Display | |
| Data in CIF |  emd_36783_validation.cif.gz emd_36783_validation.cif.gz | 16.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36783 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36783 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36783 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36783 | HTTPS FTP |
-Related structure data
| Related structure data |  8k11MC  8k10C  8k12C  8k13C  8k1bC  8k1dC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_36783.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36783.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.82 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_36783_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
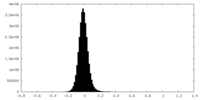
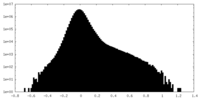
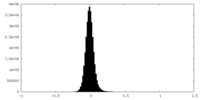
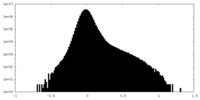
| Density Histograms |
-Half map: #1
| File | emd_36783_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : human SID1 transmembrane family member 2 (SIDT2) homodimer, ECD f...
| Entire | Name: human SID1 transmembrane family member 2 (SIDT2) homodimer, ECD focused refinement |
|---|---|
| Components |
|
-Supramolecule #1: human SID1 transmembrane family member 2 (SIDT2) homodimer, ECD f...
| Supramolecule | Name: human SID1 transmembrane family member 2 (SIDT2) homodimer, ECD focused refinement type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: SID1 transmembrane family member 2
| Macromolecule | Name: SID1 transmembrane family member 2 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 32.273844 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKANLLVLLC ALAAADAHLG VLGPKNVSQK DAEFERTYVD EVNSELVNIY TFNHTVTRNR TEGVRVSVNV LNKQKGAPLL FVVRQKEAV VSFQVPLILR GMFQRKYLYQ KVERTLCQPP TKNESEIQFF YVDVSTLSPV NTTYQLRVSR MDDFVLRTGE Q FSFNTTAA ...String: MKANLLVLLC ALAAADAHLG VLGPKNVSQK DAEFERTYVD EVNSELVNIY TFNHTVTRNR TEGVRVSVNV LNKQKGAPLL FVVRQKEAV VSFQVPLILR GMFQRKYLYQ KVERTLCQPP TKNESEIQFF YVDVSTLSPV NTTYQLRVSR MDDFVLRTGE Q FSFNTTAA QPQYFKYEFP EGVDSVIVKV TSNKAFPCSV ISIQDVLCPV YDLDNNVAFI GMYQTMTKKA AITVQRKDFP SN SFYVVVV VKTEDQACGG SLPFYPFAED EPVDQGHRQK TLSVLVSQA UniProtKB: SID1 transmembrane family member 2 |
-Macromolecule #3: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 3 / Number of copies: 8 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 281 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.2 µm |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.3 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 187510 |
| Initial angle assignment | Type: COMMON LINE |
| Final angle assignment | Type: COMMON LINE |
 Movie
Movie Controller
Controller






 Z
Z Y
Y X
X