+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | membrane proteins | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | enzyme / acetylation / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationheparan-alpha-glucosaminide N-acetyltransferase / heparan-alpha-glucosaminide N-acetyltransferase activity / MPS IIIC - Sanfilippo syndrome C / heparan sulfate proteoglycan catabolic process / HS-GAG degradation / lysosomal transport / acyltransferase activity / tertiary granule membrane / specific granule membrane / lysosomal lumen ...heparan-alpha-glucosaminide N-acetyltransferase / heparan-alpha-glucosaminide N-acetyltransferase activity / MPS IIIC - Sanfilippo syndrome C / heparan sulfate proteoglycan catabolic process / HS-GAG degradation / lysosomal transport / acyltransferase activity / tertiary granule membrane / specific granule membrane / lysosomal lumen / protein complex oligomerization / lysosomal membrane / Neutrophil degranulation / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
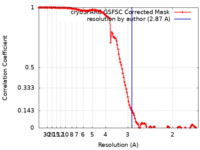
| Method | single particle reconstruction / cryo EM / Resolution: 2.87 Å | |||||||||
 Authors Authors | Yu J / Ge JP / Ruisheng X | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: membrane proteins Authors: Yu J / Ge JP / Ruisheng X | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36376.map.gz emd_36376.map.gz | 229.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36376-v30.xml emd-36376-v30.xml emd-36376.xml emd-36376.xml | 14.9 KB 14.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_36376_fsc.xml emd_36376_fsc.xml | 13.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_36376.png emd_36376.png | 92.7 KB | ||
| Filedesc metadata |  emd-36376.cif.gz emd-36376.cif.gz | 5.7 KB | ||
| Others |  emd_36376_half_map_1.map.gz emd_36376_half_map_1.map.gz emd_36376_half_map_2.map.gz emd_36376_half_map_2.map.gz | 226.5 MB 226.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36376 http://ftp.pdbj.org/pub/emdb/structures/EMD-36376 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36376 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36376 | HTTPS FTP |
-Validation report
| Summary document |  emd_36376_validation.pdf.gz emd_36376_validation.pdf.gz | 764.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_36376_full_validation.pdf.gz emd_36376_full_validation.pdf.gz | 764.3 KB | Display | |
| Data in XML |  emd_36376_validation.xml.gz emd_36376_validation.xml.gz | 21.6 KB | Display | |
| Data in CIF |  emd_36376_validation.cif.gz emd_36376_validation.cif.gz | 27.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36376 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36376 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36376 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-36376 | HTTPS FTP |
-Related structure data
| Related structure data |  8jkvMC  8jl1C  8jl3C  8jl4C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_36376.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36376.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_36376_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_36376_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : HG
| Entire | Name: HG |
|---|---|
| Components |
|
-Supramolecule #1: HG
| Supramolecule | Name: HG / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Heparan-alpha-glucosaminide N-acetyltransferase
| Macromolecule | Name: Heparan-alpha-glucosaminide N-acetyltransferase / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: heparan-alpha-glucosaminide N-acetyltransferase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 77.183992 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MTGARASAAE QRRAGRSGQA RAAERAAGMS GAGRALAALL LAASVLSAAL LAPGGSSGRD AQAAPPRDLD KKRHAELKMD QALLLIHNE LLWTNLTVYW KSECCYHCLF QVLVNVPQSP KAGKPSAAAA SVSTQHGSIL QLNDTLEEKE VCRLEYRFGE F GNYSLLVK ...String: MTGARASAAE QRRAGRSGQA RAAERAAGMS GAGRALAALL LAASVLSAAL LAPGGSSGRD AQAAPPRDLD KKRHAELKMD QALLLIHNE LLWTNLTVYW KSECCYHCLF QVLVNVPQSP KAGKPSAAAA SVSTQHGSIL QLNDTLEEKE VCRLEYRFGE F GNYSLLVK NIHNGVSEIA CDLAVNEDPV DSNLPVSIAF LIGLAVIIVI SFLRLLLSLD DFNNWISKAI SSRETDRLIN SE LGSPSRT DPLDGDVQPA TWRLSALPPR LRSVDTFRGI ALILMVFVNY GGGKYWYFKH ASWNGLTVAD LVFPWFVFIM GSS IFLSMT SILQRGCSKF RLLGKIAWRS FLLICIGIII VNPNYCLGPL SWDKVRIPGV LQRLGVTYFV VAVLELLFAK PVPE HCASE RSCLSLRDIT SSWPQWLLIL VLEGLWLGLT FLLPVPGCPT GYLGPGGIGD FGKYPNCTGG AAGYIDRLLL GDDHL YQHP SSAVLYHTEV AYDPEGILGT INSIVMAFLG VQAGKILLYY KARTKDILIR FTAWCCILGL ISVALTKVSE NEGFIP VNK NLWSLSYVTT LSSFAFFILL VLYPVVDVKG LWTGTPFFYP GMNSILVYVG HEVFENYFPF QWKLKDNQSH KEHLTQN IV ATALWVLIAY ILYRKKIFWK IGSGGGGSGG GGSGGGWSHP QFEKGGGSGG GSGGSAWSHP QFEK UniProtKB: Heparan-alpha-glucosaminide N-acetyltransferase |
-Macromolecule #3: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 3 / Number of copies: 2 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #4: DODECANE
| Macromolecule | Name: DODECANE / type: ligand / ID: 4 / Number of copies: 6 / Formula: D12 |
|---|---|
| Molecular weight | Theoretical: 170.335 Da |
| Chemical component information |  ChemComp-D12: |
-Macromolecule #5: N-OCTANE
| Macromolecule | Name: N-OCTANE / type: ligand / ID: 5 / Number of copies: 30 / Formula: OCT |
|---|---|
| Molecular weight | Theoretical: 114.229 Da |
| Chemical component information |  ChemComp-OCT: |
-Macromolecule #6: TETRADECANE
| Macromolecule | Name: TETRADECANE / type: ligand / ID: 6 / Number of copies: 24 / Formula: C14 |
|---|---|
| Molecular weight | Theoretical: 198.388 Da |
| Chemical component information |  ChemComp-C14: |
-Macromolecule #7: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 7 / Number of copies: 2 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Macromolecule #8: DECANE
| Macromolecule | Name: DECANE / type: ligand / ID: 8 / Number of copies: 2 / Formula: D10 |
|---|---|
| Molecular weight | Theoretical: 142.282 Da |
| Chemical component information |  ChemComp-D10: |
-Macromolecule #9: HEXADECANE
| Macromolecule | Name: HEXADECANE / type: ligand / ID: 9 / Number of copies: 2 / Formula: R16 |
|---|---|
| Molecular weight | Theoretical: 226.441 Da |
| Chemical component information |  ChemComp-R16: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z
Z Y
Y X
X