+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryo-EM structure of PGE1-bound hMRP4 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | cryo-EM structure of PGE1-bound hMRP4 / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationkynurenic acid metabolic process / 15-hydroxyprostaglandin dehydrogenase (NAD+) activity / purine nucleotide transmembrane transporter activity / cAMP transport / guanine nucleotide transmembrane transporter activity / ABC-type bile acid transporter activity / platelet dense granule membrane / L-tryptophan metabolic process / platelet degranulation / leukotriene transport ...kynurenic acid metabolic process / 15-hydroxyprostaglandin dehydrogenase (NAD+) activity / purine nucleotide transmembrane transporter activity / cAMP transport / guanine nucleotide transmembrane transporter activity / ABC-type bile acid transporter activity / platelet dense granule membrane / L-tryptophan metabolic process / platelet degranulation / leukotriene transport / urate transport / prostaglandin transport / glutathione transmembrane transporter activity / ABC-type glutathione-S-conjugate transporter / prostaglandin transmembrane transporter activity / ABC-type glutathione S-conjugate transporter activity / urate transmembrane transporter activity / : / external side of apical plasma membrane / prostaglandin secretion / xenobiotic transmembrane transport / export across plasma membrane / Paracetamol ADME / ABC-type xenobiotic transporter / Azathioprine ADME / Translocases; Catalysing the translocation of other compounds; Linked to the hydrolysis of a nucleoside triphosphate / ABC-type xenobiotic transporter activity / bile acid and bile salt transport / ATPase-coupled transmembrane transporter activity / efflux transmembrane transporter activity / cilium assembly / xenobiotic transmembrane transporter activity / ABC-type transporter activity / transport across blood-brain barrier / xenobiotic metabolic process / ABC-family protein mediated transport / transmembrane transport / Platelet degranulation / basolateral plasma membrane / apical plasma membrane / nucleolus / Golgi apparatus / ATP hydrolysis activity / ATP binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.95 Å | |||||||||
 Authors Authors | Liu ZM / Huang Y | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2023 Journal: Commun Biol / Year: 2023Title: Structural basis for substrate and inhibitor recognition of human multidrug transporter MRP4. Authors: Ying Huang / Chenyang Xue / Liangdong Wang / Ruiqian Bu / Jianqiang Mu / Yong Wang / Zhongmin Liu /  Abstract: Human multidrug resistance protein 4 (hMRP4, also known as ABCC4), with a representative topology of the MRP subfamily, translocates various substrates across the membrane and contributes to the ...Human multidrug resistance protein 4 (hMRP4, also known as ABCC4), with a representative topology of the MRP subfamily, translocates various substrates across the membrane and contributes to the development of multidrug resistance. However, the underlying transport mechanism of hMRP4 remains unclear due to a lack of high-resolution structures. Here, we use cryogenic electron microscopy (cryo-EM) to resolve its near-atomic structures in the apo inward-open and the ATP-bound outward-open states. We also capture the PGE1 substrate-bound structure and, importantly, the inhibitor-bound structure of hMRP4 in complex with sulindac, revealing that substrate and inhibitor compete for the same hydrophobic binding pocket although with different binding modes. Moreover, our cryo-EM structures, together with molecular dynamics simulations and biochemical assay, shed light on the structural basis of the substrate transport and inhibition mechanism, with implications for the development of hMRP4-targeted drugs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35836.map.gz emd_35836.map.gz | 97 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35836-v30.xml emd-35836-v30.xml emd-35836.xml emd-35836.xml | 16.8 KB 16.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_35836.png emd_35836.png | 37.3 KB | ||
| Filedesc metadata |  emd-35836.cif.gz emd-35836.cif.gz | 6.7 KB | ||
| Others |  emd_35836_half_map_1.map.gz emd_35836_half_map_1.map.gz emd_35836_half_map_2.map.gz emd_35836_half_map_2.map.gz | 95.5 MB 95.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35836 http://ftp.pdbj.org/pub/emdb/structures/EMD-35836 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35836 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35836 | HTTPS FTP |
-Related structure data
| Related structure data |  8iz9MC  8iz7C  8iz8C  8izaC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35836.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35836.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.855 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_35836_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_35836_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : PGE1-bound hMRP4
| Entire | Name: PGE1-bound hMRP4 |
|---|---|
| Components |
|
-Supramolecule #1: PGE1-bound hMRP4
| Supramolecule | Name: PGE1-bound hMRP4 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: ATP-binding cassette sub-family C member 4
| Macromolecule | Name: ATP-binding cassette sub-family C member 4 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO EC number: Translocases; Catalysing the translocation of other compounds; Linked to the hydrolysis of a nucleoside triphosphate |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 153.171359 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MLPVYQEVKP NPLQDANLCS RVFFWWLNPL FKIGHKRRLE EDDMYSVLPE DRSQHLGEEL QGFWDKEVLR AENDAQKPSL TRAIIKCYW KSYLVLGIFT LIEESAKVIQ PIFLGKIINY FENYDPMDSV ALNTAYAYAT VLTFCTLILA ILHHLYFYHV Q CAGMRLRV ...String: MLPVYQEVKP NPLQDANLCS RVFFWWLNPL FKIGHKRRLE EDDMYSVLPE DRSQHLGEEL QGFWDKEVLR AENDAQKPSL TRAIIKCYW KSYLVLGIFT LIEESAKVIQ PIFLGKIINY FENYDPMDSV ALNTAYAYAT VLTFCTLILA ILHHLYFYHV Q CAGMRLRV AMCHMIYRKA LRLSNMAMGK TTTGQIVNLL SNDVNKFDQV TVFLHFLWAG PLQAIAVTAL LWMEIGISCL AG MAVLIIL LPLQSCFGKL FSSLRSKTAT FTDARIRTMN EVITGIRIIK MYAWEKSFSN LITNLRKKEI SKILRSSCLR GMN LASFFS ASKIIVFVTF TTYVLLGSVI TASRVFVAVT LYGAVRLTVT LFFPSAIERV SEAIVSIRRI QTFLLLDEIS QRNR QLPSD GKKMVHVQDF TAFWDKASET PTLQGLSFTV RPGELLAVVG PVGAGKSSLL SAVLGELAPS HGLVSVHGRI AYVSQ QPWV FSGTLRSNIL FGKKYEKERY EKVIKACALK KDLQLLEDGD LTVIGDRGTT LSGGQKARVN LARAVYQDAD IYLLDD PLS AVDAEVSRHL FELCICQILH EKITILVTHQ LQYLKAASQI LILKDGKMVQ KGTYTEFLKS GIDFGSLLKK DNEESEQ PP VPGTPTLRNR TFSESSVWSQ QSSRPSLKDG ALESQDTENV PVTLSEENRS EGKVGFQAYK NYFRAGAHWI VFIFLILL N TAAQVAYVLQ DWWLSYWANK QSMLNVTVNG GGNVTEKLDL NWYLGIYSGL TVATVLFGIA RSLLVFYVLV NSSQTLHNK MFESILKAPV LFFDRNPIGR ILNRFSKDIG HLDDLLPLTF LDFIQTLLQV VGVVSVAVAV IPWIAIPLVP LGIIFIFLRR YFLETSRDV KRLESTTRSP VFSHLSSSLQ GLWTIRAYKA EERCQELFDA HQDLHSEAWF LFLTTSRWFA VRLDAICAMF V IIVAFGSL ILAKTLDAGQ VGLALSYALT LMGMFQWCVR QSAEVENMMI SVERVIEYTD LEKEAPWEYQ KRPPPAWPHE GV IIFDNVN FMYSPGGPLV LKHLTALIKS QEKVGIVGRT GAGKSSLISA LFRLSEPEGK IWIDKILTTE IGLHDLRKKM SII PQEPVL FTGTMRKNLD PFNEHTDEEL WNALQEVQLK ETIEDLPGKM DTELAESGSN FSVGQRQLVC LARAILRKNQ ILII DEATA NVDPRTDELI QKKIREKFAH CTVLTIAHRL NTIIDSDKIM VLDSGRLKEY DEPYVLLQNK ESLFYKMVQQ LGKAE AAAL TETAKQVYFK RNYPHIGHTD HMVTNTSNGQ PSTLTIFETA LLEGGGSGGG SDYKDHDGDY KDHDIDYKDD DDK UniProtKB: ATP-binding cassette sub-family C member 4 |
-Macromolecule #2: 7-[(1R,3R)-3-hydroxy-2-[(1E,3S)-3-hydroxyoct-1-en-1-yl]-5-oxocycl...
| Macromolecule | Name: 7-[(1R,3R)-3-hydroxy-2-[(1E,3S)-3-hydroxyoct-1-en-1-yl]-5-oxocyclopentyl]heptanoic acid type: ligand / ID: 2 / Number of copies: 1 / Formula: XPG |
|---|---|
| Molecular weight | Theoretical: 354.481 Da |
| Chemical component information | 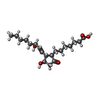 ChemComp-XPG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)