+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
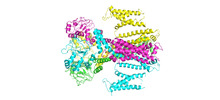
| Title | Herg1a-herg1b closed state 2 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | channelrhodopsin / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationmembrane repolarization during action potential / inward rectifier potassium channel activity / regulation of ventricular cardiac muscle cell membrane repolarization / monoatomic ion channel complex / regulation of heart rate by cardiac conduction Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Zhang MF | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structure of Herg1a-herg1b open state Authors: Zhang MF | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35608.map.gz emd_35608.map.gz | 229.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35608-v30.xml emd-35608-v30.xml emd-35608.xml emd-35608.xml | 12.9 KB 12.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_35608.png emd_35608.png | 44.8 KB | ||
| Filedesc metadata |  emd-35608.cif.gz emd-35608.cif.gz | 5.5 KB | ||
| Others |  emd_35608_half_map_1.map.gz emd_35608_half_map_1.map.gz emd_35608_half_map_2.map.gz emd_35608_half_map_2.map.gz | 225.8 MB 225.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35608 http://ftp.pdbj.org/pub/emdb/structures/EMD-35608 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35608 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35608 | HTTPS FTP |
-Validation report
| Summary document |  emd_35608_validation.pdf.gz emd_35608_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_35608_full_validation.pdf.gz emd_35608_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_35608_validation.xml.gz emd_35608_validation.xml.gz | 15.8 KB | Display | |
| Data in CIF |  emd_35608_validation.cif.gz emd_35608_validation.cif.gz | 18.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35608 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35608 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35608 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35608 | HTTPS FTP |
-Related structure data
| Related structure data |  8io5MC  8io4C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35608.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35608.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.849 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_35608_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
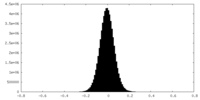
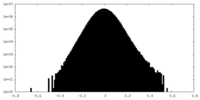
| Density Histograms |
-Half map: #2
| File | emd_35608_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
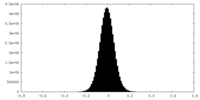
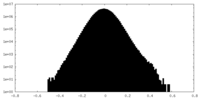
| Density Histograms |
- Sample components
Sample components
-Entire : Herg1a-herg1b channel
| Entire | Name: Herg1a-herg1b channel |
|---|---|
| Components |
|
-Supramolecule #1: Herg1a-herg1b channel
| Supramolecule | Name: Herg1a-herg1b channel / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Potassium voltage-gated channel subfamily H member 2
| Macromolecule | Name: Potassium voltage-gated channel subfamily H member 2 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 126.802727 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MPVRRGHVAP QNTFLDTIIR KFEGQSRKFI IANARVENCA VIYCNDGFCE LCGYSRAEVM QRPCTCDFLH GPRTQRRAAA QIAQALLGA EERKVEIAFY RKDGSCFLCL VDVVPVKNED GAVIMFILNF EVVMEKDMVG SPAHDTNHRG PPTSWLAPGR A KTFRLKLP ...String: MPVRRGHVAP QNTFLDTIIR KFEGQSRKFI IANARVENCA VIYCNDGFCE LCGYSRAEVM QRPCTCDFLH GPRTQRRAAA QIAQALLGA EERKVEIAFY RKDGSCFLCL VDVVPVKNED GAVIMFILNF EVVMEKDMVG SPAHDTNHRG PPTSWLAPGR A KTFRLKLP ALLALTARES SVRSGGAGGA GAPGAVVVDV DLTPAAPSSE SLALDEVTAM DNHVAGLGPA EERRALVGPG SP PRSAPGQ LPSPRAHSLN PDASGSSCSL ARTRSRESCA SVRRASSADD IEAMRAGVLP PPPRHASTGA MHPLRSGLLN STS DSDLVR YRTISKIPQI TLNFVDLKGD PFLASPTSDR EIIAPKIKER THNVTEKVTQ VLSLGADVLP EYKLQAPRIH RWTI LHYSP FKAVWDWLIL LLVIYTAVFT PYSAAFLLKE TEEGPPATEC GYACQPLAVV DLIVDIMFIV DILINFRTTY VNANE EVVS HPGRIAVHYF KGWFLIDMVA AIPFDLLIFG SGSEELIGLL KTARLLRLVR VARKLDRYSE YGAAVLFLLM CTFALI AHW LACIWYAIGN MEQPHMDSRI GWLHNLGDQI GKPYNSSGLG GPSIKDKYVT ALYFTFSSLT SVGFGNVSPN TNSEKIF SI CVMLIGSLMY ASIFGNVSAI IQRLYSGTAR YHTQMLRVRE FIRFHQIPNP LRQRLEEYFQ HAWSYTNGID MNAVLKGF P ECLQADICLH LNRSLLQHCK PFRGATKGCL RALAMKFKTT HAPPGDTLVH AGDLLTALYF ISRGSIEILR GDVVVAILG KNDIFGEPLN LYARPGKSNG DVRALTYCDL HKIHRDDLLE VLDMYPEFSD HFWSSLEITF NLRDTNMIPG SPGSTELEGG FSRQRKRKL SFRRRTDKDT EQPGEVSALG PGRAGAGPSS RGRPGGPWGE SPSSGPSSPE SSEDEGPGRS SSPLRLVPFS S PRPPGEPP GGEPLMEDCE KSSDTCNPLS GAFSGVSNIF SFWGDSRGRQ YQELPRCPAP TPSLLNIPLS SPGRRPRGDV ES RLDALQR QLNRLETRLS ADMATVLQLL QRQMTLVPPA YSAVTTPGPG PTSTSPLLPV SPLPTLTLDS LSQVSQFMAC EEL PPGAPE LPQEGPTRRL SLPGQLGALT SQPLHRHGSD PGS UniProtKB: Potassium voltage-gated channel subfamily H member 2 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.5 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.8 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 39000 |
| Initial angle assignment | Type: NOT APPLICABLE |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)