+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of CXCR3 complexed with antagonist SCH546738 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | G protein coupled receptor / chemokine receptor / CXCR3 / SCH546738 / complex / SIGNALING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationresponse to acrylamide / dynorphin receptor activity / regulation of saliva secretion / regulation of leukocyte migration / sensory perception of temperature stimulus / chemokine binding / positive regulation of eating behavior / C-X-C chemokine binding / adenylate cyclase-inhibiting opioid receptor signaling pathway / negative regulation of luteinizing hormone secretion ...response to acrylamide / dynorphin receptor activity / regulation of saliva secretion / regulation of leukocyte migration / sensory perception of temperature stimulus / chemokine binding / positive regulation of eating behavior / C-X-C chemokine binding / adenylate cyclase-inhibiting opioid receptor signaling pathway / negative regulation of luteinizing hormone secretion / chemokine receptor activity / C-X-C chemokine receptor activity / G protein-coupled opioid receptor activity / G protein-coupled opioid receptor signaling pathway / positive regulation of dopamine secretion / positive regulation of chemotaxis / T cell chemotaxis / C-C chemokine receptor activity / positive regulation of potassium ion transmembrane transport / receptor serine/threonine kinase binding / C-C chemokine binding / maternal behavior / negative regulation of execution phase of apoptosis / positive regulation of p38MAPK cascade / sensory perception / neuropeptide binding / Chemokine receptors bind chemokines / eating behavior / negative regulation of endothelial cell proliferation / positive regulation of execution phase of apoptosis / conditioned place preference / estrous cycle / regulation of cell adhesion / MECP2 regulates neuronal receptors and channels / behavioral response to cocaine / T-tubule / sensory perception of pain / axon terminus / negative regulation of angiogenesis / Peptide ligand-binding receptors / positive regulation of release of sequestered calcium ion into cytosol / sarcoplasmic reticulum / response to nicotine / cell chemotaxis / neuropeptide signaling pathway / locomotory behavior / calcium-mediated signaling / cellular response to glucose stimulus / electron transport chain / response to insulin / response to estrogen / chemotaxis / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / positive regulation of angiogenesis / synaptic vesicle membrane / adenylate cyclase-activating G protein-coupled receptor signaling pathway / positive regulation of cytosolic calcium ion concentration / cellular response to lipopolysaccharide / signaling receptor activity / presynaptic membrane / angiogenesis / G alpha (i) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / defense response to virus / chemical synaptic transmission / response to ethanol / perikaryon / periplasmic space / electron transfer activity / postsynaptic membrane / cell surface receptor signaling pathway / cell adhesion / neuron projection / immune response / iron ion binding / G protein-coupled receptor signaling pathway / inflammatory response / external side of plasma membrane / apoptotic process / heme binding / positive regulation of cell population proliferation / dendrite / cell surface / positive regulation of transcription by RNA polymerase II / mitochondrion / nucleoplasm / membrane / plasma membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
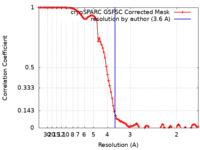
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Jiao HZ / Hu HL | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2024 Journal: Nat Struct Mol Biol / Year: 2024Title: Structural insights into the activation and inhibition of CXC chemokine receptor 3. Authors: Haizhan Jiao / Bin Pang / Aijun Liu / Qiang Chen / Qi Pan / Xiankun Wang / Yunong Xu / Ying-Chih Chiang / Ruobing Ren / Hongli Hu /  Abstract: The chemotaxis of CD4 type 1 helper cells and CD8 cytotoxic lymphocytes, guided by interferon-inducible CXC chemokine 9-11 (CXCL9-11) and CXC chemokine receptor 3 (CXCR3), plays a critical role in ...The chemotaxis of CD4 type 1 helper cells and CD8 cytotoxic lymphocytes, guided by interferon-inducible CXC chemokine 9-11 (CXCL9-11) and CXC chemokine receptor 3 (CXCR3), plays a critical role in type 1 immunity. Here we determined the structures of human CXCR3-DNG complexes activated by chemokine CXCL11, peptidomimetic agonist PS372424 and biaryl-type agonist VUF11222, and the structure of inactive CXCR3 bound to noncompetitive antagonist SCH546738. Structural analysis revealed that PS372424 shares a similar orthosteric binding pocket to the N terminus of CXCL11, while VUF11222 buries deeper and activates the receptor in a distinct manner. We showed an allosteric binding site between TM5 and TM6, accommodating SCH546738 in the inactive CXCR3. SCH546738 may restrain the receptor at an inactive state by preventing the repacking of TM5 and TM6. By revealing the binding patterns and the pharmacological properties of the four modulators, we present the activation mechanisms of CXCR3 and provide insights for future drug development. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34917.map.gz emd_34917.map.gz | 78.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34917-v30.xml emd-34917-v30.xml emd-34917.xml emd-34917.xml | 22.5 KB 22.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_34917_fsc.xml emd_34917_fsc.xml | 9.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_34917.png emd_34917.png | 88.3 KB | ||
| Filedesc metadata |  emd-34917.cif.gz emd-34917.cif.gz | 6.9 KB | ||
| Others |  emd_34917_additional_1.map.gz emd_34917_additional_1.map.gz emd_34917_half_map_1.map.gz emd_34917_half_map_1.map.gz emd_34917_half_map_2.map.gz emd_34917_half_map_2.map.gz | 40.9 MB 77.7 MB 77.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34917 http://ftp.pdbj.org/pub/emdb/structures/EMD-34917 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34917 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34917 | HTTPS FTP |
-Related structure data
| Related structure data |  8hnnMC  8hnkC  8hnlC  8hnmC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34917.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34917.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_34917_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_34917_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_34917_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CXCR3-KOR-SCH546738-Nb6
| Entire | Name: CXCR3-KOR-SCH546738-Nb6 |
|---|---|
| Components |
|
-Supramolecule #1: CXCR3-KOR-SCH546738-Nb6
| Supramolecule | Name: CXCR3-KOR-SCH546738-Nb6 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Nb6
| Macromolecule | Name: Nb6 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 18.665906 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MLLVNQSHQG FNKEHTSKMV SAIVLYVLLA AAAHSAFAMA QVQLQESGGG LVQAGESLRL SCAASGTIFR LYDMGWYRRV SGNQRELVA SITSGGSTKY GDSVKGRFTI SRDNAKNTVY LQMSSLKPED TAVYYCNAEY RTGIWEELLD GWGQGTQVTV S SHHHHHHH H |
-Macromolecule #2: Soluble cytochrome b562,C-X-C chemokine receptor type 3,Kappa-typ...
| Macromolecule | Name: Soluble cytochrome b562,C-X-C chemokine receptor type 3,Kappa-type opioid receptor type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 56.327051 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKTIIALSYI FCLVFADYKD DDDKGSADLE DNWETLNDNL KVIEKADNAA QVKDALTKMR AAALDAQKAT PPKLEDKSPD SPEMKDFRH GFDILVGQID DALKLANEGK VKEAQAAAEQ LKTTRNAYIQ KYLLVPRGSM VLEVSDHQVL NDAEVAALLE N FSSSYDYG ...String: MKTIIALSYI FCLVFADYKD DDDKGSADLE DNWETLNDNL KVIEKADNAA QVKDALTKMR AAALDAQKAT PPKLEDKSPD SPEMKDFRH GFDILVGQID DALKLANEGK VKEAQAAAEQ LKTTRNAYIQ KYLLVPRGSM VLEVSDHQVL NDAEVAALLE N FSSSYDYG ENESDSCCTS PPCPQDFSLN FDRAFLPALY SLLFLLGLLG NGAVAAVLLS RRTALSSTDT FLLHLAVADT LL VLTLPLW AVDAAVQWVF GSGLCKVAGA LFNINFYAGA LLLACISFDR YLNIVHATQL YRRGPPARVT LTCLAVWGLC LLF ALPDFI FLSAHHDERL NATHCQYNFP QVGRTALRVL QLVAGFLLPL LVMAYCYAHI LARLKSVRLL SGSREKDRNL RRIT RLVVV VVVAFALCWT PYHLVVLVDI LMDLGALARN CGRESRVDVA KSVTSGLGYM HCCLNPLLYA FVGVKFRERM WMLLL RLGC PNQRGLQRQP SSSRRDSSWS ETS UniProtKB: Soluble cytochrome b562, C-X-C chemokine receptor type 3, Kappa-type opioid receptor, C-X-C chemokine receptor type 3 |
-Macromolecule #3: 3-azanyl-6-chloranyl-5-[(3S)-4-[1-[(4-chlorophenyl)methyl]piperid...
| Macromolecule | Name: 3-azanyl-6-chloranyl-5-[(3S)-4-[1-[(4-chlorophenyl)methyl]piperidin-4-yl]-3-ethyl-piperazin-1-yl]pyrazine-2-carboxamide type: ligand / ID: 3 / Number of copies: 1 / Formula: 43I |
|---|---|
| Molecular weight | Theoretical: 492.445 Da |
| Chemical component information |  ChemComp-43I: |
-Macromolecule #4: CHOLESTEROL
| Macromolecule | Name: CHOLESTEROL / type: ligand / ID: 4 / Number of copies: 1 / Formula: CLR |
|---|---|
| Molecular weight | Theoretical: 386.654 Da |
| Chemical component information |  ChemComp-CLR: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5.0 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 53.43 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 105000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)