[English] 日本語
 Yorodumi
Yorodumi- EMDB-34730: Cryo-EM structure of SARS-CoV-2 prototype spike protein receptor-... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of SARS-CoV-2 prototype spike protein receptor-binding domain in complex with white-tailed deer ACE2 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationHydrolases; Acting on peptide bonds (peptidases) / peptidyl-dipeptidase activity / carboxypeptidase activity / metallopeptidase activity / symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region ...Hydrolases; Acting on peptide bonds (peptidases) / peptidyl-dipeptidase activity / carboxypeptidase activity / metallopeptidase activity / symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell / membrane fusion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / Attachment and Entry / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / endocytosis involved in viral entry into host cell / receptor ligand activity / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / proteolysis / : / membrane / metal ion binding / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Odocoileus virginianus texanus (mammal) Odocoileus virginianus texanus (mammal) | |||||||||
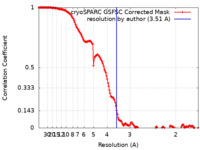
| Method | single particle reconstruction / cryo EM / Resolution: 3.51 Å | |||||||||
 Authors Authors | Han P / Meng YM / Qi JX | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: J Virol / Year: 2023 Journal: J Virol / Year: 2023Title: Structural basis of white-tailed deer, , ACE2 recognizing all the SARS-CoV-2 variants of concern with high affinity. Authors: Pu Han / Yumin Meng / Di Zhang / Zepeng Xu / Zhiyuan Li / Xiaoqian Pan / Zhennan Zhao / Linjie Li / Lingfeng Tang / Jianxun Qi / Kefang Liu / George F Gao /  Abstract: SARS-CoV-2 has been expanding its host range, among which the white-tailed deer (WTD), became the first wildlife species infected on a large scale and might serve as a host reservoir for variants of ...SARS-CoV-2 has been expanding its host range, among which the white-tailed deer (WTD), became the first wildlife species infected on a large scale and might serve as a host reservoir for variants of concern (VOCs) in case no longer circulating in humans. In this study, we comprehensively assessed the binding of the WTD angiotensin-converting enzyme 2 (ACE2) receptor to the spike (S) receptor-binding domains (RBDs) from the SARS-CoV-2 prototype (PT) strain and multiple variants. We found that WTD ACE2 could be broadly recognized by all of the tested RBDs. We further determined the complex structures of WTD ACE2 with PT, Omicron BA.1, and BA.4/5 S trimer. Detailed structural comparison revealed the important roles of RBD residues on 486, 498, and 501 sites for WTD ACE2 binding. This study deepens our understanding of the interspecies transmission mechanisms of SARS-CoV-2 and further addresses the importance of constant monitoring on SARS-CoV-2 infections in wild animals. IMPORTANCE Even if we manage to eliminate the virus among humans, it will still circulate among wildlife and continuously be transmitted back to humans. A recent study indicated that WTD may serve as reservoir for nearly extinct SARS-CoV-2 strains. Therefore, it is critical to evaluate the binding abilities of SARS-CoV-2 variants to the WTD ACE2 receptor and elucidate the molecular mechanisms of binding of the RBDs to assess the risk of spillback events. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34730.map.gz emd_34730.map.gz | 169.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34730-v30.xml emd-34730-v30.xml emd-34730.xml emd-34730.xml | 16 KB 16 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_34730_fsc.xml emd_34730_fsc.xml | 14.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_34730.png emd_34730.png | 43.5 KB | ||
| Filedesc metadata |  emd-34730.cif.gz emd-34730.cif.gz | 6.1 KB | ||
| Others |  emd_34730_half_map_1.map.gz emd_34730_half_map_1.map.gz emd_34730_half_map_2.map.gz emd_34730_half_map_2.map.gz | 318 MB 318 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34730 http://ftp.pdbj.org/pub/emdb/structures/EMD-34730 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34730 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34730 | HTTPS FTP |
-Related structure data
| Related structure data |  8hg0MC  8hfxC  8hfyC  8hfzC  8ifyC  8ifzC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34730.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34730.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_34730_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_34730_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : SARS-CoV-2 prototype spike protein receptor-binding domain in com...
| Entire | Name: SARS-CoV-2 prototype spike protein receptor-binding domain in complex with white-tailed deer ACE2 |
|---|---|
| Components |
|
-Supramolecule #1: SARS-CoV-2 prototype spike protein receptor-binding domain in com...
| Supramolecule | Name: SARS-CoV-2 prototype spike protein receptor-binding domain in complex with white-tailed deer ACE2 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Angiotensin-converting enzyme
| Macromolecule | Name: Angiotensin-converting enzyme / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: Hydrolases; Acting on peptide bonds (peptidases) |
|---|---|
| Source (natural) | Organism:  Odocoileus virginianus texanus (mammal) Odocoileus virginianus texanus (mammal) |
| Molecular weight | Theoretical: 71.420438 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MTGSFWLLLS LVAVTAAQST TEEQAKTFLE KFNHEAEDLS YQSSLASWNY NTNITDENVQ KMNEARAKWS AFYEEQSRMA KTYSLEEIQ NLTLKRQLKA LQQSGTSVLS AEKSKRLNTI LNTMSTIYST GKVLDPNTQE CLALEPGLDD IMENSRDYNR R LWAWEGWR ...String: MTGSFWLLLS LVAVTAAQST TEEQAKTFLE KFNHEAEDLS YQSSLASWNY NTNITDENVQ KMNEARAKWS AFYEEQSRMA KTYSLEEIQ NLTLKRQLKA LQQSGTSVLS AEKSKRLNTI LNTMSTIYST GKVLDPNTQE CLALEPGLDD IMENSRDYNR R LWAWEGWR AEVGKQLRPL YEEYVVLENE MARANNYEDY GDYWRGDYEV TEAGDYDYSR DQLMKDVENT FAEIKPLYEQ LH AYVRAKL MDTYPSYISP TGCLPAHLLG DMWGRFWTNL YSLTVPFKHK PSIDVTEKMK NQSWDAERIF KEAEKFFVSI SLP HMTQGF WDNSMLTEPG DGRKVVCHPT AWDLGKGDFR IKMCTKVTMD DFLTAHHEMG HIQYDMAYAA QPYLLRDGAN EGFH EAVGE IMSLSAATPH YLKALGLLEP DFYEDNETEI NFLLKQALTI VGTLPFTYML EKWRWMVFKG EIPKEQWMEK WWEMK REIV GVVEPLPHDE TYCDPACLFH VAEDYSFIRY YTRTIYQFQF HEALCKTANH EGALFKCDIS NSTEAGQRLL QMLSLG KSE PWTLALESIV GIKTMDVKPL LNYFEPLFTW LKEQNRNSFV GWSTEWTPYS UniProtKB: Angiotensin-converting enzyme |
-Macromolecule #2: Spike protein S1
| Macromolecule | Name: Spike protein S1 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 21.873496 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: TNLCPFGEVF NATRFASVYA WNRKRISNCV ADYSVLYNSA SFSTFKCYGV SPTKLNDLCF TNVYADSFVI RGDEVRQIAP GQTGKIADY NYKLPDDFTG CVIAWNSNNL DSKVGGNYNY LYRLFRKSNL KPFERDISTE IYQAGSTPCN GVEGFNCYFP L QSYGFQPT ...String: TNLCPFGEVF NATRFASVYA WNRKRISNCV ADYSVLYNSA SFSTFKCYGV SPTKLNDLCF TNVYADSFVI RGDEVRQIAP GQTGKIADY NYKLPDDFTG CVIAWNSNNL DSKVGGNYNY LYRLFRKSNL KPFERDISTE IYQAGSTPCN GVEGFNCYFP L QSYGFQPT NGVGYQPYRV VVLSFELLHA PATVCGP UniProtKB: Spike glycoprotein |
-Macromolecule #3: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 3 / Number of copies: 3 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #4: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 4 / Number of copies: 1 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)