| 登録情報 | データベース: EMDB / ID: EMD-34478
|
|---|
| タイトル | DHA-bound FFAR4 in complex with Gs |
|---|
 マップデータ マップデータ | |
|---|
 試料 試料 | - 複合体: GPCR/G-protein complex
- タンパク質・ペプチド: engineered mini Galpha-S subunit
- タンパク質・ペプチド: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
- タンパク質・ペプチド: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
- タンパク質・ペプチド: Nb35
- タンパク質・ペプチド: Free fatty acid receptor 4
- リガンド: DOCOSA-4,7,10,13,16,19-HEXAENOIC ACID
|
|---|
 キーワード キーワード | GPCR-G-protein complex / MEMBRANE PROTEIN-IMMUNE SYSTEM complex |
|---|
| 機能・相同性 |  機能・相同性情報 機能・相同性情報
negative regulation of somatostatin secretion / positive regulation of glucagon secretion / ghrelin secretion / Free fatty acid receptors / regulation of D-glucose transmembrane transport / taste receptor activity / hormone secretion / arrestin family protein binding / negative regulation of cytokine production / negative regulation of interleukin-1 beta production ...negative regulation of somatostatin secretion / positive regulation of glucagon secretion / ghrelin secretion / Free fatty acid receptors / regulation of D-glucose transmembrane transport / taste receptor activity / hormone secretion / arrestin family protein binding / negative regulation of cytokine production / negative regulation of interleukin-1 beta production / ciliary membrane / white fat cell differentiation / endocytic vesicle / positive regulation of osteoblast differentiation / brown fat cell differentiation / positive regulation of brown fat cell differentiation / fatty acid binding / G protein-coupled receptor activity / negative regulation of inflammatory response / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / G protein-coupled acetylcholine receptor signaling pathway / adenylate cyclase-activating G protein-coupled receptor signaling pathway / G-protein activation / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through CDC42 / Glucagon signaling in metabolic regulation / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / photoreceptor disc membrane / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / G alpha (z) signalling events / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / ADORA2B mediated anti-inflammatory cytokines production / cellular response to catecholamine stimulus / ADP signalling through P2Y purinoceptor 1 / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / Inactivation, recovery and regulation of the phototransduction cascade / cellular response to prostaglandin E stimulus / G-protein beta-subunit binding / heterotrimeric G-protein complex / G alpha (12/13) signalling events / sensory perception of taste / extracellular vesicle / signaling receptor complex adaptor activity / Thrombin signalling through proteinase activated receptors (PARs) / positive regulation of cold-induced thermogenesis / GTPase binding / positive regulation of cytosolic calcium ion concentration / Ca2+ pathway / retina development in camera-type eye / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / fibroblast proliferation / G alpha (i) signalling events / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (q) signalling events / cilium / ciliary basal body / Ras protein signal transduction / Extra-nuclear estrogen signaling / cell population proliferation / positive regulation of ERK1 and ERK2 cascade / endosome membrane / G protein-coupled receptor signaling pathway / inflammatory response / lysosomal membrane / intracellular membrane-bounded organelle / GTPase activity / synapse / protein-containing complex binding / negative regulation of apoptotic process / signal transduction / extracellular exosome / membrane / plasma membrane / cytosol / cytoplasm類似検索 - 分子機能 G-protein, gamma subunit / G-protein gamma subunit domain profile. / G-protein gamma-like domain / G-protein gamma-like domain superfamily / GGL domain / G protein gamma subunit-like motifs / GGL domain / Guanine nucleotide-binding protein, beta subunit / G-protein, beta subunit / G protein-coupled receptor, rhodopsin-like ...G-protein, gamma subunit / G-protein gamma subunit domain profile. / G-protein gamma-like domain / G-protein gamma-like domain superfamily / GGL domain / G protein gamma subunit-like motifs / GGL domain / Guanine nucleotide-binding protein, beta subunit / G-protein, beta subunit / G protein-coupled receptor, rhodopsin-like / GPCR, rhodopsin-like, 7TM / G-protein coupled receptors family 1 profile. / 7 transmembrane receptor (rhodopsin family) / G-protein beta WD-40 repeat / WD40 repeat, conserved site / Trp-Asp (WD) repeats signature. / WD domain, G-beta repeat / Trp-Asp (WD) repeats profile. / Trp-Asp (WD) repeats circular profile. / WD40 repeats / WD40 repeat / WD40-repeat-containing domain superfamily / WD40/YVTN repeat-like-containing domain superfamily類似検索 - ドメイン・相同性 Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 / Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 / Free fatty acid receptor 4類似検索 - 構成要素 |
|---|
| 生物種 |  Homo sapiens (ヒト) / Homo sapiens (ヒト) /   Lama glama (ラマ) Lama glama (ラマ) |
|---|
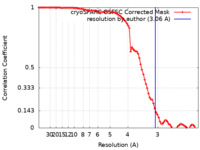
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.06 Å |
|---|
 データ登録者 データ登録者 | He Y / Yin H |
|---|
| 資金援助 |  中国, 1件 中国, 1件 | Organization | Grant number | 国 |
|---|
| National Natural Science Foundation of China (NSFC) | 32070048 |  中国 中国 |
|
|---|
 引用 引用 |  ジャーナル: Cell Res / 年: 2023 ジャーナル: Cell Res / 年: 2023
タイトル: Structural basis of omega-3 fatty acid receptor FFAR4 activation and G protein coupling selectivity.
著者: Han Yin / Asuka Inoue / Zhengxiong Ma / Xinyan Zhu / Ruixue Xia / Zhenmei Xu / Na Wang / Yaning Duan / Anqi Zhang / Changyou Guo / Yuanzheng He /   |
|---|
| 履歴 | | 登録 | 2022年10月10日 | - |
|---|
| ヘッダ(付随情報) 公開 | 2023年6月21日 | - |
|---|
| マップ公開 | 2023年6月21日 | - |
|---|
| 更新 | 2024年11月20日 | - |
|---|
| 現状 | 2024年11月20日 | 処理サイト: RCSB / 状態: 公開 |
|---|
|
|---|
 データを開く
データを開く 基本情報
基本情報
 マップデータ
マップデータ 試料
試料 キーワード
キーワード 機能・相同性情報
機能・相同性情報 Homo sapiens (ヒト) /
Homo sapiens (ヒト) / 
 データ登録者
データ登録者 中国, 1件
中国, 1件  引用
引用 ジャーナル: Cell Res / 年: 2023
ジャーナル: Cell Res / 年: 2023

 構造の表示
構造の表示 ダウンロードとリンク
ダウンロードとリンク emd_34478.map.gz
emd_34478.map.gz EMDBマップデータ形式
EMDBマップデータ形式 emd-34478-v30.xml
emd-34478-v30.xml emd-34478.xml
emd-34478.xml EMDBヘッダ
EMDBヘッダ emd_34478_fsc.xml
emd_34478_fsc.xml FSCデータファイル
FSCデータファイル emd_34478.png
emd_34478.png emd-34478.cif.gz
emd-34478.cif.gz emd_34478_half_map_1.map.gz
emd_34478_half_map_1.map.gz emd_34478_half_map_2.map.gz
emd_34478_half_map_2.map.gz http://ftp.pdbj.org/pub/emdb/structures/EMD-34478
http://ftp.pdbj.org/pub/emdb/structures/EMD-34478 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34478
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34478 emd_34478_validation.pdf.gz
emd_34478_validation.pdf.gz EMDB検証レポート
EMDB検証レポート emd_34478_full_validation.pdf.gz
emd_34478_full_validation.pdf.gz emd_34478_validation.xml.gz
emd_34478_validation.xml.gz emd_34478_validation.cif.gz
emd_34478_validation.cif.gz https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34478
https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34478 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34478
ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34478 リンク
リンク EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource マップ
マップ ダウンロード / ファイル: emd_34478.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
ダウンロード / ファイル: emd_34478.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) 試料の構成要素
試料の構成要素 Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト)
 Homo sapiens (ヒト)
Homo sapiens (ヒト)
 Homo sapiens (ヒト)
Homo sapiens (ヒト)


 Homo sapiens (ヒト)
Homo sapiens (ヒト)

 解析
解析 試料調製
試料調製 電子顕微鏡法
電子顕微鏡法 FIELD EMISSION GUN
FIELD EMISSION GUN
 ムービー
ムービー コントローラー
コントローラー
































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)