+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | apo-ferritin embedded in crystalline ice frozen at -183 degree | |||||||||
 Map data Map data | Apo-ferritin map from crystal-embedded datasets frozen at -183 degree collected via FEI Arctica by K2 camera | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationiron ion sequestering activity / ferritin complex / Scavenging by Class A Receptors / Golgi Associated Vesicle Biogenesis / ferroxidase / autolysosome / negative regulation of ferroptosis / ferroxidase activity / negative regulation of fibroblast proliferation / ferric iron binding ...iron ion sequestering activity / ferritin complex / Scavenging by Class A Receptors / Golgi Associated Vesicle Biogenesis / ferroxidase / autolysosome / negative regulation of ferroptosis / ferroxidase activity / negative regulation of fibroblast proliferation / ferric iron binding / autophagosome / iron ion transport / Iron uptake and transport / ferrous iron binding / tertiary granule lumen / ficolin-1-rich granule lumen / intracellular iron ion homeostasis / immune response / iron ion binding / negative regulation of cell population proliferation / Neutrophil degranulation / extracellular exosome / extracellular region / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / Resolution: 2.4 Å | |||||||||
 Authors Authors | Zhang X / Shi H / Wu C | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2023 Journal: Structure / Year: 2023Title: Addressing compressive deformation of proteins embedded in crystalline ice. Authors: Huigang Shi / Chunling Wu / Xinzheng Zhang /  Abstract: For cryoelectron microscopy (cryo-EM), high cooling rates have been required for preparation of protein samples to vitrify the surrounding water and avoid formation of damaging crystalline ice. ...For cryoelectron microscopy (cryo-EM), high cooling rates have been required for preparation of protein samples to vitrify the surrounding water and avoid formation of damaging crystalline ice. Whether and how crystalline ice affects single-particle cryo-EM is still unclear. Here, single-particle cryo-EM was used to analyze three-dimensional structures of various proteins and viruses embedded in crystalline ice formed at various cooling rates. Low cooling rates led to shrinkage deformation and density distortions on samples having loose structures. Higher cooling rates reduced deformations. Deformation-free proteins in crystalline ice were obtained by modifying the freezing conditions, and reconstructions from these samples revealed a marked improvement over vitreous ice. This procedure also increased the efficiency of cryo-EM structure determinations and was essential for high-resolution reconstructions. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_32703.map.gz emd_32703.map.gz | 95.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-32703-v30.xml emd-32703-v30.xml emd-32703.xml emd-32703.xml | 23 KB 23 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_32703_fsc.xml emd_32703_fsc.xml | 13.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_32703.png emd_32703.png | 138.3 KB | ||
| Masks |  emd_32703_msk_1.map emd_32703_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Others |  emd_32703_additional_1.map.gz emd_32703_additional_1.map.gz emd_32703_additional_2.map.gz emd_32703_additional_2.map.gz emd_32703_additional_3.map.gz emd_32703_additional_3.map.gz emd_32703_additional_4.map.gz emd_32703_additional_4.map.gz emd_32703_additional_5.map.gz emd_32703_additional_5.map.gz emd_32703_half_map_1.map.gz emd_32703_half_map_1.map.gz emd_32703_half_map_2.map.gz emd_32703_half_map_2.map.gz | 201.5 MB 59.9 MB 59.8 MB 59.8 MB 59.7 MB 188.2 MB 188.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-32703 http://ftp.pdbj.org/pub/emdb/structures/EMD-32703 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32703 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32703 | HTTPS FTP |
-Related structure data
| Related structure data |  8f4lM  8hhsMC  8bqnC  8ew0C  8ew2C  8f49C  8f7yC  8hi2C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_32703.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_32703.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Apo-ferritin map from crystal-embedded datasets frozen at -183 degree collected via FEI Arctica by K2 camera | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.5 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_32703_msk_1.map emd_32703_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
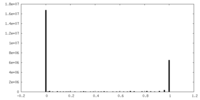
| Density Histograms |
-Additional map: Apo-ferritin map from standard freezing datasets collected via...
| File | emd_32703_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Apo-ferritin map from standard freezing datasets collected via FEI Arctica by K2 camera | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Apo-ferritin map from crystal-embedded datasets frozen at -140...
| File | emd_32703_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Apo-ferritin map from crystal-embedded datasets frozen at -140 degree collected via Titan Krios by K2 camera | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Apo-ferritin map from standard freezing datasets collected via...
| File | emd_32703_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Apo-ferritin map from standard freezing datasets collected via Titan Krios by K2 camera | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Apo-ferritin map from crystal-embedded datasets frozen at -60...
| File | emd_32703_additional_4.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Apo-ferritin map from crystal-embedded datasets frozen at -60 degree collected via Titan Krios by K2 camera | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Apo-ferritin map from crystal-embedded datasets frozen at -100...
| File | emd_32703_additional_5.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Apo-ferritin map from crystal-embedded datasets frozen at -100 degree collected via Titan Krios by K2 camera | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Apo-ferritin map from crystal-embedded datasets frozen at -183...
| File | emd_32703_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Apo-ferritin map from crystal-embedded datasets frozen at -183 degree collected via FEI Arctica by K2 camera. The half map of apo-ferritin was flipped and centered. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Apo-ferritin map from crystal-embedded datasets frozen at -183...
| File | emd_32703_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Apo-ferritin map from crystal-embedded datasets frozen at -183 degree collected via FEI Arctica by K2 camera. The half map of apo-ferritin was flipped and centered. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human apo-ferritin
| Entire | Name: Human apo-ferritin |
|---|---|
| Components |
|
-Supramolecule #1: Human apo-ferritin
| Supramolecule | Name: Human apo-ferritin / type: complex / ID: 1 / Chimera: Yes / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 440 KDa |
-Experimental details
-Structure determination
 Processing Processing | single particle reconstruction |
|---|---|
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Sugar embedding | Material: crystalline ice |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)