[English] 日本語
 Yorodumi
Yorodumi- EMDB-31223: FOOT AND MOUTH DISEASE VIRUS O/TIBET/99-BOUND THE SINGLE CHAIN FR... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-31223 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | FOOT AND MOUTH DISEASE VIRUS O/TIBET/99-BOUND THE SINGLE CHAIN FRAGMEN ANTIBODY C4 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | FOOT AND MOUTH DISEASE VIRUS / FMDV / VIRUS | |||||||||
| Function / homology |  Function and homology information Function and homology informationimmunoglobulin complex / adaptive immune response / extracellular region / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Foot-and-mouth disease virus / Foot-and-mouth disease virus /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.75 Å | |||||||||
 Authors Authors | He Y / Li K | |||||||||
 Citation Citation |  Journal: J Virol / Year: 2021 Journal: J Virol / Year: 2021Title: Two Cross-Protective Antigen Sites on Foot-and-Mouth Disease Virus Serotype O Structurally Revealed by Broadly Neutralizing Antibodies from Cattle. Authors: Kun Li / Yong He / Li Wang / Pinghua Li / Sheng Wang / Pu Sun / Huifang Bao / Yimei Cao / Xuerong Liu / Guoqiang Zhu / Yali Song / Xingwen Bai / Xueqing Ma / Yuanfang Fu / Hong Yuan / Jing ...Authors: Kun Li / Yong He / Li Wang / Pinghua Li / Sheng Wang / Pu Sun / Huifang Bao / Yimei Cao / Xuerong Liu / Guoqiang Zhu / Yali Song / Xingwen Bai / Xueqing Ma / Yuanfang Fu / Hong Yuan / Jing Zhang / Jian Wang / Yingli Chen / Dong Li / Zhiyong Lou / Zaixin Liu / Zengjun Lu /  Abstract: Foot-and-mouth disease virus (FMDV) is a highly contagious virus that infects cloven-hoofed animals. Neutralizing antibodies play critical roles in antiviral infection. Although five known antigen ...Foot-and-mouth disease virus (FMDV) is a highly contagious virus that infects cloven-hoofed animals. Neutralizing antibodies play critical roles in antiviral infection. Although five known antigen sites that induce neutralizing antibodies have been defined, studies on cross-protective antigen sites are still scarce. We mapped two cross-protective antigen sites using 13 bovine-derived broadly neutralizing monoclonal antibodies (bnAbs) capable of neutralizing 4 lineages within 3 topotypes of FMDV serotype O. One antigen site was formed by a novel cluster of VP3-focused epitopes recognized by bnAb C4 and C4-like antibodies. The cryo-electron microscopy (cryo-EM) structure of the FMDV-OTi (O/Tibet/99)-C4 complex showed close contact with VP3 and a novel interprotomer antigen epitope around the icosahedral 3-fold axis of the FMDV particle, which is far beyond the known antigen site 4. The key determinants of the neutralizing function of C4 and C4-like antibodies on the capsid were βB (T65), the B-C loop (T68), the E-F loop (E131 and K134), and the H-I loop (G196), revealing a novel antigen site on VP3. The other antigen site comprised two group epitopes on VP2 recognized by 9 bnAbs (B57, B73, B77, B82, F28, F145, F150, E46, and E54), which belong to the known antigen site 2 of FMDV serotype O. Notably, bnAb C4 potently promoted FMDV RNA release in response to damage to viral particles, suggesting that the targeted epitope contains a trigger mechanism for particle disassembly. This study revealed two cross-protective antigen sites that can elicit cross-reactive neutralizing antibodies in cattle and provided new structural information for the design of a broad-spectrum molecular vaccine against FMDV serotype O. FMDV is the causative agent of foot-and-mouth disease (FMD), which is one of the most contagious and economically devastating diseases of domestic animals. The antigenic structure of FMDV serotype O is rather complicated, especially for those sites that can elicit a cross-protective neutralizing antibody response. Monoclonal neutralization antibodies provide both crucial defense components against FMDV infection and valuable tools for fine analysis of the antigenic structure. In this study, we found a cluster of novel VP3-focused epitopes using 13 bnAbs against FMDV serotype O from natural host cattle, which revealed two cross-protective antigen sites on VP2 and VP3. Antibody C4 targeting this novel epitope potently promoted viral particle disassembly and RNA release before infection, which may indicate a vulnerable region of FMDV. This study reveals new structural information about cross-protective antigen sites of FMDV serotype O, providing valuable and strong support for future research on broad-spectrum vaccines against FMD. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_31223.map.gz emd_31223.map.gz | 96.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-31223-v30.xml emd-31223-v30.xml emd-31223.xml emd-31223.xml | 15.7 KB 15.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_31223.png emd_31223.png | 284.7 KB | ||
| Filedesc metadata |  emd-31223.cif.gz emd-31223.cif.gz | 5.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-31223 http://ftp.pdbj.org/pub/emdb/structures/EMD-31223 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31223 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-31223 | HTTPS FTP |
-Related structure data
| Related structure data |  7eo0MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_31223.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_31223.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.93 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
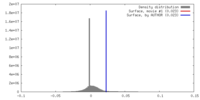
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : FMDV-OTi-C4
| Entire | Name: FMDV-OTi-C4 |
|---|---|
| Components |
|
-Supramolecule #1: FMDV-OTi-C4
| Supramolecule | Name: FMDV-OTi-C4 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Supramolecule #2: FOOT AND MOUTH DISEASE VIRUS O/TIBET/99
| Supramolecule | Name: FOOT AND MOUTH DISEASE VIRUS O/TIBET/99 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:   Foot-and-mouth disease virus Foot-and-mouth disease virus |
-Supramolecule #3: C4 scFv
| Supramolecule | Name: C4 scFv / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #5-#6 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: O/TIBET/99 VP1
| Macromolecule | Name: O/TIBET/99 VP1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Foot-and-mouth disease virus Foot-and-mouth disease virus |
| Molecular weight | Theoretical: 23.524781 KDa |
| Sequence | String: TTSTGESADP VTATVENYGG ETQVQRRQHT DVSFILDRFV KVTPKDQINV LDLMQTPAHT LVGALLRTAT YYFADLEVAV KHEGNLTWV PNGAPETALD NTTNPTAYHK APLTRLALPY TAPHRVLATV YNGNCKYGES PVTNARGDLQ VLAQKAARAL P TSFNYGAI ...String: TTSTGESADP VTATVENYGG ETQVQRRQHT DVSFILDRFV KVTPKDQINV LDLMQTPAHT LVGALLRTAT YYFADLEVAV KHEGNLTWV PNGAPETALD NTTNPTAYHK APLTRLALPY TAPHRVLATV YNGNCKYGES PVTNARGDLQ VLAQKAARAL P TSFNYGAI KATRVTELLY RMKRAETYCP RPLLAIHPSE ARHKQKIVAP VKQLL |
-Macromolecule #2: O/TIBET/99 VP2
| Macromolecule | Name: O/TIBET/99 VP2 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Foot-and-mouth disease virus Foot-and-mouth disease virus |
| Molecular weight | Theoretical: 24.338387 KDa |
| Sequence | String: DKKTEETTLL EDRILTTRNG HTTSTTQSSV GVTYGYATAE DFVSGPNTSG LETRVVQAER FFKTHLFDWV TSDPFGRCYQ LELPTDHKG VYGSLTDSYA YMRNGWDVEV TAVGNQFNGG CLLVAMVPEL CSIDKRGLYQ LTLFPHQFIN PRTNMTAHIT V PFVGVNRY ...String: DKKTEETTLL EDRILTTRNG HTTSTTQSSV GVTYGYATAE DFVSGPNTSG LETRVVQAER FFKTHLFDWV TSDPFGRCYQ LELPTDHKG VYGSLTDSYA YMRNGWDVEV TAVGNQFNGG CLLVAMVPEL CSIDKRGLYQ LTLFPHQFIN PRTNMTAHIT V PFVGVNRY DQYKVHKPWT LVVMVVAPLT VNTEGAPQIK VYANIAPTNV HVAGEFPSKE |
-Macromolecule #3: O/TIBET/99 VP3
| Macromolecule | Name: O/TIBET/99 VP3 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Foot-and-mouth disease virus Foot-and-mouth disease virus |
| Molecular weight | Theoretical: 23.875801 KDa |
| Sequence | String: GIFPVACSDG YGGLVTTDPK TADPAYGKVF NPPRNMLPGR FTNFLDVAEA CPTFLHFEGD VPYVTTKTDS DRVLAQFDLS LAAKHMSNT FLAGLAQYYT QYSGTINLHF MFTGPTDAKA RYMIAYAPPG MEPPKTPEAA AHCIHAEWDT GLNSKFTFSI P YLSAADYA ...String: GIFPVACSDG YGGLVTTDPK TADPAYGKVF NPPRNMLPGR FTNFLDVAEA CPTFLHFEGD VPYVTTKTDS DRVLAQFDLS LAAKHMSNT FLAGLAQYYT QYSGTINLHF MFTGPTDAKA RYMIAYAPPG MEPPKTPEAA AHCIHAEWDT GLNSKFTFSI P YLSAADYA YTASDAAETT NVQGWVCLFQ ITHGKADGDA LVVLASAGKD FELRLPVDAR TQ |
-Macromolecule #4: O/TIBET/99 VP4
| Macromolecule | Name: O/TIBET/99 VP4 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Foot-and-mouth disease virus Foot-and-mouth disease virus |
| Molecular weight | Theoretical: 8.778129 KDa |
| Sequence | String: GAGQSSPATG SQNQSGNTGS IINNYYMQQY QNSMDTQLGD NAISGGSNEG STDTTSTHTT NTQNNDWFSK LASSAFSGLF GALLA |
-Macromolecule #5: Ig heavy chain variable region
| Macromolecule | Name: Ig heavy chain variable region / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.914535 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: QVQLRESGPS LVKPSQTLFL TCTVSGFSLT SYSVNWVRQT PGKMLECLGG IATSGSTGYN PVLKSRLRIT KDNSKSQVSL SVSNVTPED TATYYCAKWS SRGGYDCGVH SSDYSYLDAW GQGLLVTVSS UniProtKB: Ig heavy chain variable region |
-Macromolecule #6: Ig lamda chain variable region
| Macromolecule | Name: Ig lamda chain variable region / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 12.79488 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: WAQAVLTQPS SVSASLGQRV SITCSGSSSN IGRYGATWYQ QVPGSGLRTI IYGSSRRPSG VPDRFSGSKS GNTVTLTISS LQPEDEADY FCAAYDISTN AVFGSGTTLT LLGDYKDDDD KGG UniProtKB: Ig lamda chain variable region |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: OTHER |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: EMDB MAP |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.75 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 14458 |
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)