[English] 日本語
 Yorodumi
Yorodumi- EMDB-29562: Cryo-EM structure of Cas1:Cas2-DEDDh:PAM-containing prespacer complex -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Cas1:Cas2-DEDDh:PAM-containing prespacer complex | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | CRISPR / integrase / CRISPR adaptation module / PAM / prespacer / exonuclease / DNA binding protein-DNA complex / enzyme / ribonucleoprotein | ||||||||||||
| Biological species |  Megasphaera (bacteria) Megasphaera (bacteria) | ||||||||||||
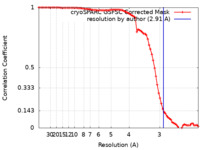
| Method | single particle reconstruction / cryo EM / Resolution: 2.91 Å | ||||||||||||
 Authors Authors | Skopintsev P / Tuck OT / Soczek KM / Doudna J | ||||||||||||
| Funding support |  United States, United States,  Switzerland, 3 items Switzerland, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nature / Year: 2023 Journal: Nature / Year: 2023Title: Genome expansion by a CRISPR trimmer-integrase. Authors: Joy Y Wang / Owen T Tuck / Petr Skopintsev / Katarzyna M Soczek / Gary Li / Basem Al-Shayeb / Julia Zhou / Jennifer A Doudna /  Abstract: CRISPR-Cas adaptive immune systems capture DNA fragments from invading mobile genetic elements and integrate them into the host genome to provide a template for RNA-guided immunity. CRISPR systems ...CRISPR-Cas adaptive immune systems capture DNA fragments from invading mobile genetic elements and integrate them into the host genome to provide a template for RNA-guided immunity. CRISPR systems maintain genome integrity and avoid autoimmunity by distinguishing between self and non-self, a process for which the CRISPR/Cas1-Cas2 integrase is necessary but not sufficient. In some microorganisms, the Cas4 endonuclease assists CRISPR adaptation, but many CRISPR-Cas systems lack Cas4. Here we show here that an elegant alternative pathway in a type I-E system uses an internal DnaQ-like exonuclease (DEDDh) to select and process DNA for integration using the protospacer adjacent motif (PAM). The natural Cas1-Cas2/exonuclease fusion (trimmer-integrase) catalyses coordinated DNA capture, trimming and integration. Five cryo-electron microscopy structures of the CRISPR trimmer-integrase, visualized both before and during DNA integration, show how asymmetric processing generates size-defined, PAM-containing substrates. Before genome integration, the PAM sequence is released by Cas1 and cleaved by the exonuclease, marking inserted DNA as self and preventing aberrant CRISPR targeting of the host. Together, these data support a model in which CRISPR systems lacking Cas4 use fused or recruited exonucleases for faithful acquisition of new CRISPR immune sequences. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29562.map.gz emd_29562.map.gz | 40.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29562-v30.xml emd-29562-v30.xml emd-29562.xml emd-29562.xml | 22.1 KB 22.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_29562_fsc.xml emd_29562_fsc.xml | 7.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_29562.png emd_29562.png | 95.2 KB | ||
| Masks |  emd_29562_msk_1.map emd_29562_msk_1.map | 52.7 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-29562.cif.gz emd-29562.cif.gz | 6.3 KB | ||
| Others |  emd_29562_additional_1.map.gz emd_29562_additional_1.map.gz emd_29562_half_map_1.map.gz emd_29562_half_map_1.map.gz emd_29562_half_map_2.map.gz emd_29562_half_map_2.map.gz | 25.7 MB 39.8 MB 39.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29562 http://ftp.pdbj.org/pub/emdb/structures/EMD-29562 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29562 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29562 | HTTPS FTP |
-Related structure data
| Related structure data |  8fyaMC  8fy9C  8fybC  8fycC  8fydC C: citing same article ( M: atomic model generated by this map |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_29562.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29562.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.115 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_29562_msk_1.map emd_29562_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_29562_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_29562_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_29562_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cas1:Cas2-DEDDh:PAM-containing prespacer complex
| Entire | Name: Cas1:Cas2-DEDDh:PAM-containing prespacer complex |
|---|---|
| Components |
|
-Supramolecule #1: Cas1:Cas2-DEDDh:PAM-containing prespacer complex
| Supramolecule | Name: Cas1:Cas2-DEDDh:PAM-containing prespacer complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Megasphaera (bacteria) Megasphaera (bacteria) |
| Molecular weight | Theoretical: 224 KDa |
-Macromolecule #1: Cas2-DEDDh
| Macromolecule | Name: Cas2-DEDDh / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Megasphaera (bacteria) Megasphaera (bacteria) |
| Molecular weight | Theoretical: 32.632615 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPMTVITLKN VPQSLRGDLT RWMQEIATGV YVGNFNSRIR EYLWRRVQET MGAGEASMCF AARNELGYDF LTENASRSVI DYDGLPLIF IPKEQSAVSD LPKGFSTAAK LHRAHIAGSG KKKEKPIRYV VIDIETDGKD AKRNHILEIG AIRCEDGKET H FTALISGD ...String: MPMTVITLKN VPQSLRGDLT RWMQEIATGV YVGNFNSRIR EYLWRRVQET MGAGEASMCF AARNELGYDF LTENASRSVI DYDGLPLIF IPKEQSAVSD LPKGFSTAAK LHRAHIAGSG KKKEKPIRYV VIDIETDGKD AKRNHILEIG AIRCEDGKET H FTALISGD AVPPSITKLT GITATLLQKE GQEEKKVLTA FREFIGDDDL VGYHVSFDIE FLRQAFKKYG LGYLKNKTHD LL RIVKKEQ LFQADYKLET SLQSYGIHKK VPHRALGDAE LVKCLAKKLN KF |
-Macromolecule #2: Cas1
| Macromolecule | Name: Cas1 / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Megasphaera (bacteria) Megasphaera (bacteria) |
| Molecular weight | Theoretical: 35.022074 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAGPIIAGKS ESSELPRVED RATFIYIEHA KINRVDSAVT VAEAKGVVRI PAAMIGVLLL GPGTDISHRA VELLGDTGTA LVWVGEQGV RYYASGRALA RSTRFLVKQA ELVTNERSRL RVARRMYQMR FPTEDVSKLT MQQLRSHEGA RVRRKYRELS K KYNVPWKK ...String: MAGPIIAGKS ESSELPRVED RATFIYIEHA KINRVDSAVT VAEAKGVVRI PAAMIGVLLL GPGTDISHRA VELLGDTGTA LVWVGEQGV RYYASGRALA RSTRFLVKQA ELVTNERSRL RVARRMYQMR FPTEDVSKLT MQQLRSHEGA RVRRKYRELS K KYNVPWKK RVYNPDDFAG GDPINQALSA AHVALYGLVH SVVAALGLSP GLGFVHTGHD RSFIYDVADL YKAEITVPIA FA VAAEAEE GQDIGQLARL RTRDAFVDGK ILKRMVKDLQ TLLEIPEEGQ IEAEPLSLWD DKEKLVPYGV NYSEVTSCP |
-Macromolecule #3: DNA (28-MER)
| Macromolecule | Name: DNA (28-MER) / type: dna / ID: 3 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Megasphaera (bacteria) Megasphaera (bacteria) |
| Molecular weight | Theoretical: 8.629578 KDa |
| Sequence | String: (DG)(DC)(DA)(DA)(DC)(DC)(DA)(DC)(DT)(DT) (DG)(DT)(DG)(DC)(DA)(DT)(DC)(DA)(DT)(DG) (DA)(DG)(DT)(DG)(DA)(DT)(DG)(DA) |
-Macromolecule #4: DNA (31-MER)
| Macromolecule | Name: DNA (31-MER) / type: dna / ID: 4 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Megasphaera (bacteria) Megasphaera (bacteria) |
| Molecular weight | Theoretical: 10.137508 KDa |
| Sequence | String: (DA)(DC)(DT)(DC)(DA)(DT)(DG)(DA)(DT)(DG) (DC)(DA)(DC)(DA)(DA)(DG)(DT)(DG)(DG)(DT) (DT)(DG)(DC)(DG)(DC)(DG)(DT)(DG)(DT) (DT)(DC)(DC)(DC) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 281 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.0 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER |
|---|---|
| Output model |  PDB-8fya: |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)