+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Structure of Agrobacterium tumefaciens bacteriophage Milano curved tail | |||||||||
 マップデータ マップデータ | ||||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | Myophage / redox trigger / VIRUS | |||||||||
| 機能・相同性 | Tail sheath protein / Virion-associated protein 機能・相同性情報 機能・相同性情報 | |||||||||
| 生物種 |  Agrobacterium phage Milano (ファージ) Agrobacterium phage Milano (ファージ) | |||||||||
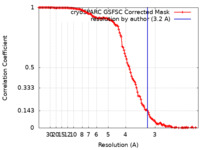
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.2 Å | |||||||||
 データ登録者 データ登録者 | Sonani RR / Leiman PG / Wang F / Kreutzberger MAB / Sebastian A / Esteves NC / Kelly RJ / Scharf B / Egelman EH | |||||||||
| 資金援助 |  米国, 1件 米国, 1件
| |||||||||
 引用 引用 |  ジャーナル: Nat Commun / 年: 2024 ジャーナル: Nat Commun / 年: 2024タイトル: An extensive disulfide bond network prevents tail contraction in Agrobacterium tumefaciens phage Milano. 著者: Ravi R Sonani / Lee K Palmer / Nathaniel C Esteves / Abigail A Horton / Amanda L Sebastian / Rebecca J Kelly / Fengbin Wang / Mark A B Kreutzberger / William K Russell / Petr G Leiman / ...著者: Ravi R Sonani / Lee K Palmer / Nathaniel C Esteves / Abigail A Horton / Amanda L Sebastian / Rebecca J Kelly / Fengbin Wang / Mark A B Kreutzberger / William K Russell / Petr G Leiman / Birgit E Scharf / Edward H Egelman /  要旨: A contractile sheath and rigid tube assembly is a widespread apparatus used by bacteriophages, tailocins, and the bacterial type VI secretion system to penetrate cell membranes. In this mechanism, ...A contractile sheath and rigid tube assembly is a widespread apparatus used by bacteriophages, tailocins, and the bacterial type VI secretion system to penetrate cell membranes. In this mechanism, contraction of an external sheath powers the motion of an inner tube through the membrane. The structure, energetics, and mechanism of the machinery imply rigidity and straightness. The contractile tail of Agrobacterium tumefaciens bacteriophage Milano is flexible and bent to varying degrees, which sets it apart from other contractile tail-like systems. Here, we report structures of the Milano tail including the sheath-tube complex, baseplate, and putative receptor-binding proteins. The flexible-to-rigid transformation of the Milano tail upon contraction can be explained by unique electrostatic properties of the tail tube and sheath. All components of the Milano tail, including sheath subunits, are crosslinked by disulfides, some of which must be reduced for contraction to occur. The putative receptor-binding complex of Milano contains a tailspike, a tail fiber, and at least two small proteins that form a garland around the distal ends of the tailspikes and tail fibers. Despite being flagellotropic, Milano lacks thread-like tail filaments that can wrap around the flagellum, and is thus likely to employ a different binding mechanism. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_29353.map.gz emd_29353.map.gz | 203.9 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-29353-v30.xml emd-29353-v30.xml emd-29353.xml emd-29353.xml | 15 KB 15 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_29353_fsc.xml emd_29353_fsc.xml | 12.7 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_29353.png emd_29353.png | 60.3 KB | ||
| Filedesc metadata |  emd-29353.cif.gz emd-29353.cif.gz | 5.5 KB | ||
| その他 |  emd_29353_half_map_1.map.gz emd_29353_half_map_1.map.gz emd_29353_half_map_2.map.gz emd_29353_half_map_2.map.gz | 200.7 MB 200.7 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29353 http://ftp.pdbj.org/pub/emdb/structures/EMD-29353 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29353 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29353 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_29353.map.gz / 形式: CCP4 / 大きさ: 216 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_29353.map.gz / 形式: CCP4 / 大きさ: 216 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-ハーフマップ: #1
| ファイル | emd_29353_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: #2
| ファイル | emd_29353_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : Agrobacterium phage Milano
| 全体 | 名称:  Agrobacterium phage Milano (ファージ) Agrobacterium phage Milano (ファージ) |
|---|---|
| 要素 |
|
-超分子 #1: Agrobacterium phage Milano
| 超分子 | 名称: Agrobacterium phage Milano / タイプ: virus / ID: 1 / 親要素: 0 / 含まれる分子: all / NCBI-ID: 2557550 / 生物種: Agrobacterium phage Milano / ウイルスタイプ: VIRION / ウイルス・単離状態: SPECIES / ウイルス・エンベロープ: Yes / ウイルス・中空状態: No |
|---|---|
| 宿主 | 生物種:  Agrobacterium fabrum str. C58 (バクテリア) Agrobacterium fabrum str. C58 (バクテリア) |
-分子 #1: Virion-associated protein
| 分子 | 名称: Virion-associated protein / タイプ: protein_or_peptide / ID: 1 / コピー数: 18 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Agrobacterium phage Milano (ファージ) Agrobacterium phage Milano (ファージ) |
| 分子量 | 理論値: 14.673427 KDa |
| 配列 | 文字列: MACNKQNGVK NILITFTDCD TQEVIGPISH EQPDDTLPTY KNCAWTNTAL TNGYVQRSAS NATMTLPVVR DLRVPLAFYQ GCAQVDVQV EKFDGTVMTL TEGAVVEPEE SDGRSVTMNI VASEIDELLP PGSLAAA UniProtKB: Virion-associated protein |
-分子 #2: Tail sheath protein
| 分子 | 名称: Tail sheath protein / タイプ: protein_or_peptide / ID: 2 / コピー数: 12 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Agrobacterium phage Milano (ファージ) Agrobacterium phage Milano (ファージ) |
| 分子量 | 理論値: 53.896094 KDa |
| 配列 | 文字列: MAQDALSDGF VRLCIDPSLN FFGEGCKILV EGQMTDDGSA TPDAVTCVTS ELDIIERFGQ GSVLTESLRK VFCTCKSGVS VYALPREDA AAGVKAVYTL TIAGPATTDG RVQLYMGEAE YAVDIGVDAG DTATDIAAAI VAAISPDFPY AATAAAGVIT L TARNAGTI ...文字列: MAQDALSDGF VRLCIDPSLN FFGEGCKILV EGQMTDDGSA TPDAVTCVTS ELDIIERFGQ GSVLTESLRK VFCTCKSGVS VYALPREDA AAGVKAVYTL TIAGPATTDG RVQLYMGEAE YAVDIGVDAG DTATDIAAAI VAAISPDFPY AATAAAGVIT L TARNAGTI GNHLSVIYTN LGSCTSVTPE GVTVTFAQTT AGSVNPTPND YATVVNECCF AVYVLSSDDT DWQENLRDWI RS AWDCSKP QCFGHGYVFN KGTLGQVLAD GDNSAELSRL ALPTTYPVLP YLTNAAYGAL SACSTCNNPE LNIQGQTFGL LSC INMPES CTPGWTFGEV TQLQANGFVV SGPSTTSGQG NYTSPYIYND VTNYLRDEKN RPNATFRDAS SRRLAAATGV ALAE FLQQF NGLAVFTKNT NIRTGIIGTN PRLMLGKIRK WAQDNVGTLF SEFDNINEDI QLLTDFEVQP KCVGQPGIFH LNMRY RPPV RGARINVNMA PALFDNCDR UniProtKB: Tail sheath protein |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.4 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | TFS KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K3 (6k x 4k) / 平均電子線量: 50.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 2.2 µm / 最小 デフォーカス(公称値): 1.2 µm |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)