+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Mojiang virus F ectodomain in prefusion form | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | F ectodomain / prefusion / VIRAL PROTEIN | |||||||||
| Function / homology | Precursor fusion glycoprotein F0, Paramyxoviridae / Fusion glycoprotein F0 / fusion of virus membrane with host plasma membrane / viral envelope / host cell plasma membrane / virion membrane / plasma membrane / Fusion glycoprotein F0 Function and homology information Function and homology information | |||||||||
| Biological species |  Mojiang virus Mojiang virus | |||||||||
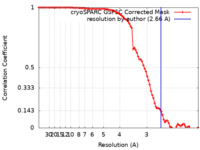
| Method | single particle reconstruction / cryo EM / Resolution: 2.66 Å | |||||||||
 Authors Authors | Low YS / Isaacs A / Modhiran N / Watterson D | |||||||||
| Funding support |  Australia, 1 items Australia, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structure and antigenicity of divergent Henipavirus fusion glycoproteins. Authors: Ariel Isaacs / Yu Shang Low / Kyle L Macauslane / Joy Seitanidou / Cassandra L Pegg / Stacey T M Cheung / Benjamin Liang / Connor A P Scott / Michael J Landsberg / Benjamin L Schulz / Keith ...Authors: Ariel Isaacs / Yu Shang Low / Kyle L Macauslane / Joy Seitanidou / Cassandra L Pegg / Stacey T M Cheung / Benjamin Liang / Connor A P Scott / Michael J Landsberg / Benjamin L Schulz / Keith J Chappell / Naphak Modhiran / Daniel Watterson /  Abstract: In August 2022, a novel henipavirus (HNV) named Langya virus (LayV) was isolated from patients with severe pneumonic disease in China. This virus is closely related to Mòjiāng virus (MojV), and ...In August 2022, a novel henipavirus (HNV) named Langya virus (LayV) was isolated from patients with severe pneumonic disease in China. This virus is closely related to Mòjiāng virus (MojV), and both are divergent from the bat-borne HNV members, Nipah (NiV) and Hendra (HeV) viruses. The spillover of LayV is the first instance of a HNV zoonosis to humans outside of NiV and HeV, highlighting the continuing threat this genus poses to human health. In this work, we determine the prefusion structures of MojV and LayV F proteins via cryogenic electron microscopy to 2.66 and 3.37 Å, respectively. We show that despite sequence divergence from NiV, the F proteins adopt an overall similar structure but are antigenically distinct as they do not react to known antibodies or sera. Glycoproteomic analysis revealed that while LayV F is less glycosylated than NiV F, it contains a glycan that shields a site of vulnerability previously identified for NiV. These findings explain the distinct antigenic profile of LayV and MojV F, despite the extent to which they are otherwise structurally similar to NiV. Our results carry implications for broad-spectrum HNV vaccines and therapeutics, and indicate an antigenic, yet not structural, divergence from prototypical HNVs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29300.map.gz emd_29300.map.gz | 32.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29300-v30.xml emd-29300-v30.xml emd-29300.xml emd-29300.xml | 19.4 KB 19.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_29300_fsc.xml emd_29300_fsc.xml | 7.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_29300.png emd_29300.png | 38.3 KB | ||
| Masks |  emd_29300_msk_1.map emd_29300_msk_1.map | 64 MB |  Mask map Mask map | |
| Others |  emd_29300_additional_1.map.gz emd_29300_additional_1.map.gz emd_29300_half_map_1.map.gz emd_29300_half_map_1.map.gz emd_29300_half_map_2.map.gz emd_29300_half_map_2.map.gz | 59.6 MB 59.5 MB 59.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29300 http://ftp.pdbj.org/pub/emdb/structures/EMD-29300 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29300 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29300 | HTTPS FTP |
-Validation report
| Summary document |  emd_29300_validation.pdf.gz emd_29300_validation.pdf.gz | 942.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_29300_full_validation.pdf.gz emd_29300_full_validation.pdf.gz | 942.4 KB | Display | |
| Data in XML |  emd_29300_validation.xml.gz emd_29300_validation.xml.gz | 15.1 KB | Display | |
| Data in CIF |  emd_29300_validation.cif.gz emd_29300_validation.cif.gz | 20.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29300 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29300 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29300 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29300 | HTTPS FTP |
-Related structure data
| Related structure data |  8fmyMC  8fmxC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_29300.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29300.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_29300_msk_1.map emd_29300_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_29300_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_29300_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_29300_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Mojiang virus F glycoprotein ectodomain in the prefusion form
| Entire | Name: Mojiang virus F glycoprotein ectodomain in the prefusion form |
|---|---|
| Components |
|
-Supramolecule #1: Mojiang virus F glycoprotein ectodomain in the prefusion form
| Supramolecule | Name: Mojiang virus F glycoprotein ectodomain in the prefusion form type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Mojiang virus Mojiang virus |
| Molecular weight | Theoretical: 200 KDa |
-Macromolecule #1: Fusion glycoprotein F0
| Macromolecule | Name: Fusion glycoprotein F0 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Mojiang virus Mojiang virus |
| Molecular weight | Theoretical: 52.636922 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MALNKNMFSS LFLGYLLVYA TTVQSSIHYD SLSKVGVIKG LTYNYKIKGS PSTKLMVVKL IPNIDSVKNC TQKQYDEYKN LVRKALEPV KMAIDTMLNN VKSGNNKYRF AGAIMAGVAL GVATAATVTA GIALHRSNEN AQAIANMKSA IQNTNEAVKQ L QLANKQTL ...String: MALNKNMFSS LFLGYLLVYA TTVQSSIHYD SLSKVGVIKG LTYNYKIKGS PSTKLMVVKL IPNIDSVKNC TQKQYDEYKN LVRKALEPV KMAIDTMLNN VKSGNNKYRF AGAIMAGVAL GVATAATVTA GIALHRSNEN AQAIANMKSA IQNTNEAVKQ L QLANKQTL AVIDTIRGEI NNNIIPVINQ LSCDTIGLSV GIRLTQYYSE IITAFGPALQ NPVNTRITIQ AISSVFNGNF DE LLKIMGY TSGDLYEILH SELIRGNIID VDVDAGYIAL EIEFPNLTLV PNAVVQELMP ISYNIDGDEW VTLVPRFVLT RTT LLSNID TSRCTITDSS VICDNDYALP MSHELIGCLQ GDTSKCAREK VVSSYVPKFA LSDGLVYANC LNTICRCMDT DTPI SQSLG ATVSLLDNKR CSVYQVGDVL ISVGSYLGDG EYNADNVELG PPIVIDKIDI GNQLAGINQT LQEAEDYIEK SEEFL KG |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.7 mg/mL |
|---|---|
| Buffer | pH: 6.8 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Instrument: LEICA EM GP |
| Details | 0.255% CHAPS added |
- Electron microscopy
Electron microscopy
| Microscope | JEOL CRYO ARM 300 |
|---|---|
| Specialist optics | Energy filter - Name: In-column Omega Filter / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 25.0 µm / Nominal defocus min: 5.0 µm / Nominal magnification: 100000 |
| Sample stage | Specimen holder model: JEOL CRYOSPECPORTER / Cooling holder cryogen: NITROGEN |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Overall B value: 102.7 |
|---|---|
| Output model |  PDB-8fmy: |
 Movie
Movie Controller
Controller







 Z
Z Y
Y X
X