+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Wildtype rat TRPV2 in nanodiscs bound to RR | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TRPV2 / TRPV5 / TRP Channel / Ruthenium Red / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationgrowth cone membrane / TRP channels / response to temperature stimulus / positive regulation of calcium ion import / calcium ion import across plasma membrane / positive regulation of axon extension / monoatomic cation channel activity / axonal growth cone / endomembrane system / calcium channel activity ...growth cone membrane / TRP channels / response to temperature stimulus / positive regulation of calcium ion import / calcium ion import across plasma membrane / positive regulation of axon extension / monoatomic cation channel activity / axonal growth cone / endomembrane system / calcium channel activity / melanosome / lamellipodium / positive regulation of cold-induced thermogenesis / cell body / axon / negative regulation of cell population proliferation / cell surface / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
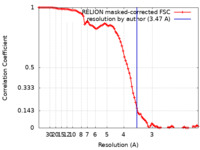
| Method | single particle reconstruction / cryo EM / Resolution: 3.47 Å | |||||||||
 Authors Authors | Pumroy RA / Protopopova AD / Rocereta JA / De Jesus-Perez JJ / Fluck EC / Moiseenkova-Bell VY | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: EMBO Rep / Year: 2024 Journal: EMBO Rep / Year: 2024Title: Molecular details of ruthenium red pore block in TRPV channels. Authors: Ruth A Pumroy / José J De Jesús-Pérez / Anna D Protopopova / Julia A Rocereta / Edwin C Fluck / Tabea Fricke / Bo-Hyun Lee / Tibor Rohacs / Andreas Leffler / Vera Moiseenkova-Bell /    Abstract: Transient receptor potential vanilloid (TRPV) channels play a critical role in calcium homeostasis, pain sensation, immunological response, and cancer progression. TRPV channels are blocked by ...Transient receptor potential vanilloid (TRPV) channels play a critical role in calcium homeostasis, pain sensation, immunological response, and cancer progression. TRPV channels are blocked by ruthenium red (RR), a universal pore blocker for a wide array of cation channels. Here we use cryo-electron microscopy to reveal the molecular details of RR block in TRPV2 and TRPV5, members of the two TRPV subfamilies. In TRPV2 activated by 2-aminoethoxydiphenyl borate, RR is tightly coordinated in the open selectivity filter, blocking ion flow and preventing channel inactivation. In TRPV5 activated by phosphatidylinositol 4,5-bisphosphate, RR blocks the selectivity filter and closes the lower gate through an interaction with polar residues in the pore vestibule. Together, our results provide a detailed understanding of TRPV subfamily pore block, the dynamic nature of the selectivity filter and allosteric communication between the selectivity filter and lower gate. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29046.map.gz emd_29046.map.gz | 59.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29046-v30.xml emd-29046-v30.xml emd-29046.xml emd-29046.xml | 14.6 KB 14.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_29046_fsc.xml emd_29046_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_29046.png emd_29046.png | 126.1 KB | ||
| Filedesc metadata |  emd-29046.cif.gz emd-29046.cif.gz | 5.8 KB | ||
| Others |  emd_29046_half_map_1.map.gz emd_29046_half_map_1.map.gz emd_29046_half_map_2.map.gz emd_29046_half_map_2.map.gz | 49.5 MB 49.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29046 http://ftp.pdbj.org/pub/emdb/structures/EMD-29046 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29046 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29046 | HTTPS FTP |
-Validation report
| Summary document |  emd_29046_validation.pdf.gz emd_29046_validation.pdf.gz | 1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_29046_full_validation.pdf.gz emd_29046_full_validation.pdf.gz | 1 MB | Display | |
| Data in XML |  emd_29046_validation.xml.gz emd_29046_validation.xml.gz | 15.9 KB | Display | |
| Data in CIF |  emd_29046_validation.cif.gz emd_29046_validation.cif.gz | 21.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29046 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29046 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29046 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29046 | HTTPS FTP |
-Related structure data
| Related structure data |  8fflMC  8ffmC  8ffnC  8ffqC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_29046.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29046.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||
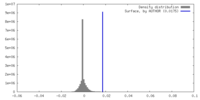
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_29046_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
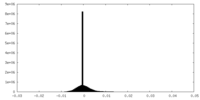
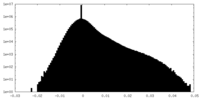
| Density Histograms |
-Half map: #1
| File | emd_29046_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Tetramer of wildtype rat TRPV2 in nanodiscs bound to ruthenium red
| Entire | Name: Tetramer of wildtype rat TRPV2 in nanodiscs bound to ruthenium red |
|---|---|
| Components |
|
-Supramolecule #1: Tetramer of wildtype rat TRPV2 in nanodiscs bound to ruthenium red
| Supramolecule | Name: Tetramer of wildtype rat TRPV2 in nanodiscs bound to ruthenium red type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 340 KDa |
-Macromolecule #1: Transient receptor potential cation channel subfamily V member 2
| Macromolecule | Name: Transient receptor potential cation channel subfamily V member 2 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 86.798891 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MTSASSPPAF RLETSDGDEE GNAEVNKGKQ EPPPMESPFQ REDRNSSPQI KVNLNFIKRP PKNTSAPSQQ EPDRFDRDRL FSVVSRGVP EELTGLLEYL RWNSKYLTDS AYTEGSTGKT CLMKAVLNLQ DGVNACIMPL LQIDKDSGNP KLLVNAQCTD E FYQGHSAL ...String: MTSASSPPAF RLETSDGDEE GNAEVNKGKQ EPPPMESPFQ REDRNSSPQI KVNLNFIKRP PKNTSAPSQQ EPDRFDRDRL FSVVSRGVP EELTGLLEYL RWNSKYLTDS AYTEGSTGKT CLMKAVLNLQ DGVNACIMPL LQIDKDSGNP KLLVNAQCTD E FYQGHSAL HIAIEKRSLQ CVKLLVENGA DVHLRACGRF FQKHQGTCFY FGELPLSLAA CTKQWDVVTY LLENPHQPAS LE ATDSLGN TVLHALVMIA DNSPENSALV IHMYDGLLQM GARLCPTVQL EEISNHQGLT PLKLAAKEGK IEIFRHILQR EFS GPYQPL SRKFTEWCYG PVRVSLYDLS SVDSWEKNSV LEIIAFHCKS PNRHRMVVLE PLNKLLQEKW DRLVSRFFFN FACY LVYMF IFTVVAYHQP SLDQPAIPSS KATFGESMLL LGHILILLGG IYLLLGQLWY FWRRRLFIWI SFMDSYFEIL FLLQA LLTV LSQVLRFMET EWYLPLLVLS LVLGWLNLLY YTRGFQHTGI YSVMIQKVIL RDLLRFLLVY LVFLFGFAVA LVSLSR EAR SPKAPEDNNS TVTEQPTVGQ EEEPAPYRSI LDASLELFKF TIGMGELAFQ EQLRFRGVVL LLLLAYVLLT YVLLLNM LI ALMSETVNHV ADNSWSIWKL QKAISVLEME NGYWWCRRKK HREGRLLKVG TRGDGTPDER WCFRVEEVNW AAWEKTLP T LSEDPSGPGI TGNKKNPTSK PGKNSASEED HLPLQVLQSP UniProtKB: Transient receptor potential cation channel subfamily V member 2 |
-Macromolecule #2: 1,2-DIDECANOYL-SN-GLYCERO-3-PHOSPHOETHANOLAMINE
| Macromolecule | Name: 1,2-DIDECANOYL-SN-GLYCERO-3-PHOSPHOETHANOLAMINE / type: ligand / ID: 2 / Number of copies: 4 / Formula: PEX |
|---|---|
| Molecular weight | Theoretical: 522.632 Da |
| Chemical component information | 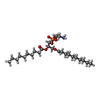 ChemComp-PEX: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 42.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 20.0 µm / Nominal defocus min: 5.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)