[English] 日本語
 Yorodumi
Yorodumi- EMDB-28603: Cryo-EM structure of cGMP bound human CNGA3/CNGB3 channel in GDN,... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of cGMP bound human CNGA3/CNGB3 channel in GDN, transition state 1 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Heterotetramer / ligand-bound / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationinorganic cation import across plasma membrane / intracellular cyclic nucleotide activated cation channel complex / intracellularly cGMP-activated cation channel activity / intracellularly cAMP-activated cation channel activity / axon initial segment / transmembrane transporter complex / myosin binding / sodium channel activity / cGMP binding / monoatomic cation transmembrane transport ...inorganic cation import across plasma membrane / intracellular cyclic nucleotide activated cation channel complex / intracellularly cGMP-activated cation channel activity / intracellularly cAMP-activated cation channel activity / axon initial segment / transmembrane transporter complex / myosin binding / sodium channel activity / cGMP binding / monoatomic cation transmembrane transport / response to magnesium ion / photoreceptor outer segment / monoatomic cation transport / response to cAMP / ligand-gated monoatomic ion channel activity / glial cell projection / visual perception / calcium channel activity / retina development in camera-type eye / perikaryon / cadherin binding / dendrite / signal transduction / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
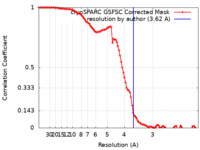
| Method | single particle reconstruction / cryo EM / Resolution: 3.62 Å | |||||||||
 Authors Authors | Hu Z / Zheng X / Yang J | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Conformational trajectory of allosteric gating of the human cone photoreceptor cyclic nucleotide-gated channel. Authors: Zhengshan Hu / Xiangdong Zheng / Jian Yang /  Abstract: Cyclic nucleotide-gated (CNG) channels transduce chemical signals into electrical signals in sensory receptors and neurons. They are activated by cGMP or cAMP, which bind to the cyclic nucleotide- ...Cyclic nucleotide-gated (CNG) channels transduce chemical signals into electrical signals in sensory receptors and neurons. They are activated by cGMP or cAMP, which bind to the cyclic nucleotide-binding domain (CNBD) to open a gate located 50-60 Å away in the central cavity. Structures of closed and open vertebrate CNG channels have been solved, but the conformational landscape of this allosteric gating remains to be elucidated and enriched. Here, we report structures of the cGMP-activated human cone photoreceptor CNGA3/CNGB3 channel in closed, intermediate, pre-open and open states in detergent or lipid nanodisc, all with fully bound cGMP. The pre-open and open states are obtained only in the lipid nanodisc, suggesting a critical role of lipids in tuning the energetic landscape of CNGA3/CNGB3 activation. The different states exhibit subunit-unique, incremental and distinct conformational rearrangements that originate in the CNBD, propagate through the gating ring to the transmembrane domain, and gradually open the S6 cavity gate. Our work illustrates a spatial conformational-change wave of allosteric gating of a vertebrate CNG channel by its natural ligand and provides an expanded framework for studying CNG properties and channelopathy. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28603.map.gz emd_28603.map.gz | 93.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28603-v30.xml emd-28603-v30.xml emd-28603.xml emd-28603.xml | 16.2 KB 16.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_28603_fsc.xml emd_28603_fsc.xml | 9.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_28603.png emd_28603.png | 195.1 KB | ||
| Filedesc metadata |  emd-28603.cif.gz emd-28603.cif.gz | 6.3 KB | ||
| Others |  emd_28603_half_map_1.map.gz emd_28603_half_map_1.map.gz emd_28603_half_map_2.map.gz emd_28603_half_map_2.map.gz | 92 MB 92 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28603 http://ftp.pdbj.org/pub/emdb/structures/EMD-28603 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28603 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28603 | HTTPS FTP |
-Related structure data
| Related structure data |  8eu3MC  8etpC  8eucC  8ev8C  8ev9C  8evaC  8evbC  8evcC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_28603.map.gz / Format: CCP4 / Size: 98.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28603.map.gz / Format: CCP4 / Size: 98.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
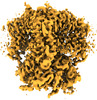
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.058 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_28603_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
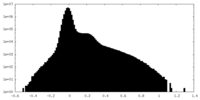
| Density Histograms |
-Half map: #1
| File | emd_28603_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : human cone photoreceptor heterotetrameric CNG channel CNGA3/CNGB3...
| Entire | Name: human cone photoreceptor heterotetrameric CNG channel CNGA3/CNGB3 in complex with cGMP in GDN |
|---|---|
| Components |
|
-Supramolecule #1: human cone photoreceptor heterotetrameric CNG channel CNGA3/CNGB3...
| Supramolecule | Name: human cone photoreceptor heterotetrameric CNG channel CNGA3/CNGB3 in complex with cGMP in GDN type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Cyclic nucleotide-gated cation channel alpha-3
| Macromolecule | Name: Cyclic nucleotide-gated cation channel alpha-3 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 78.93793 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAKINTQYSH PSRTHLKVKT SDRDLNRAEN GLSRAHSSSE ETSSVLQPGI AMETRGLADS GQGSFTGQGI ARLSRLIFLL RRWAARHVH HQDQGPDSFP DRFRGAELKE VSSQESNAQA NVGSQEPADR GRSAWPLAKC NTNTSNNTEE EKKTKKKDAI V VDPSSNLY ...String: MAKINTQYSH PSRTHLKVKT SDRDLNRAEN GLSRAHSSSE ETSSVLQPGI AMETRGLADS GQGSFTGQGI ARLSRLIFLL RRWAARHVH HQDQGPDSFP DRFRGAELKE VSSQESNAQA NVGSQEPADR GRSAWPLAKC NTNTSNNTEE EKKTKKKDAI V VDPSSNLY YRWLTAIALP VFYNWYLLIC RACFDELQSE YLMLWLVLDY SADVLYVLDV LVRARTGFLE QGLMVSDTNR LW QHYKTTT QFKLDVLSLV PTDLAYLKVG TNYPEVRFNR LLKFSRLFEF FDRTETRTNY PNMFRIGNLV LYILIIIHWN ACI YFAISK FIGFGTDSWV YPNISIPEHG RLSRKYIYSL YWSTLTLTTI GETPPPVKDE EYLFVVVDFL VGVLIFATIV GNVG SMISN MNASRAEFQA KIDSIKQYMQ FRKVTKDLET RVIRWFDYLW ANKKTVDEKE VLKSLPDKLK AEIAINVHLD TLKKV RIFQ DCEAGLLVEL VLKLRPTVFS PGDYICKKGD IGKEMYIINE GKLAVVADDG VTQFVVLSDG SYFGEISILN IKGSKS GNR RTANIRSIGY SDLFCLSKDD LMEALTEYPE AKKALEEKGR QILMKDNLID EELARAGADP KDLEEKVEQL GSSLDTL QT RFARLLAEYN ATQMKMKQRL SQLESQVKGG GDKPLADGEV PGDATKTEDK QQ UniProtKB: Cyclic nucleotide-gated channel alpha-3 |
-Macromolecule #2: Cyclic nucleotide-gated cation channel beta-3
| Macromolecule | Name: Cyclic nucleotide-gated cation channel beta-3 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 92.289492 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFKSLTKVNK VKPIGENNEN EQSSRRNEEG SHPSNQSQQT TAQEENKGEE KSLKTKSTPV TSEEPHTNIQ DKLSKKNSSG DLTTNPDPQ NAAEPTGTVP EQKEMDPGKE GPNSPQNKPP AAPVINEYAD AQLHNLVKRM RQRTALYKKK LVEGDLSSPE A SPQTAKPT ...String: MFKSLTKVNK VKPIGENNEN EQSSRRNEEG SHPSNQSQQT TAQEENKGEE KSLKTKSTPV TSEEPHTNIQ DKLSKKNSSG DLTTNPDPQ NAAEPTGTVP EQKEMDPGKE GPNSPQNKPP AAPVINEYAD AQLHNLVKRM RQRTALYKKK LVEGDLSSPE A SPQTAKPT AVPPVKESDD KPTEHYYRLL WFKVKKMPLT EYLKRIKLPN SIDSYTDRLY LLWLLLVTLA YNWNCCFIPL RL VFPYQTA DNIHYWLIAD IICDIIYLYD MLFIQPRLQF VRGGDIIVDS NELRKHYRTS TKFQLDVASI IPFDICYLFF GFN PMFRAN RMLKYTSFFE FNHHLESIMD KAYIYRVIRT TGYLLFILHI NACVYYWASN YEGIGTTRWV YDGEGNEYLR CYYW AVRTL ITIGGLPEPQ TLFEIVFQLL NFFSGVFVFS SLIGQMRDVI GAATANQNYF RACMDDTIAY MNNYSIPKLV QKRVR TWYE YTWDSQRMLD ESDLLKTLPT TVQLALAIDV NFSIISKVDL FKGCDTQMIY DMLLRLKSVL YLPGDFVCKK GEIGKE MYI IKHGEVQVLG GPDGTKVLVT LKAGSVFGEI SLLAAGGGNR RTANVVAHGF ANLLTLDKKT LQEILVHYPD SERILMK KA RVLLKQKAKT AEATPPRKDL ALLFPPKEET PKLFKTLLGG TGKASLARLL KLKREQAAQK KENSEGGEEE GKENEDKQ K ENEDKQKENE DKGKENEDKD KGREPEEKPL DRPECTASPI AVEEEPHSVR RTVLPRGTSR QSLIISMAPS AEGGEEVLT IEVKEKAKQ UniProtKB: Cyclic nucleotide-gated channel beta-3 |
-Macromolecule #3: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 3 / Number of copies: 3 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #4: CYCLIC GUANOSINE MONOPHOSPHATE
| Macromolecule | Name: CYCLIC GUANOSINE MONOPHOSPHATE / type: ligand / ID: 4 / Number of copies: 4 / Formula: PCG |
|---|---|
| Molecular weight | Theoretical: 345.205 Da |
| Chemical component information |  ChemComp-PCG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8.58 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 55.03 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: OTHER / Nominal defocus max: 1.9000000000000001 µm / Nominal defocus min: 0.9 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)