[English] 日本語
 Yorodumi
Yorodumi- EMDB-28245: PaFS prenyltransferase core with two interacting cyclase domains ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | PaFS prenyltransferase core with two interacting cyclase domains - East and North | |||||||||
 Map data Map data | PaFS prenyltransferase core with two interacting cyclase domains - East and North | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Diaporthe amygdali (fungus) Diaporthe amygdali (fungus) | |||||||||
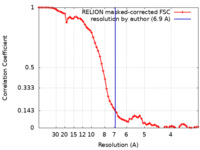
| Method | single particle reconstruction / cryo EM / Resolution: 6.9 Å | |||||||||
 Authors Authors | Faylo JL / van Eeuwen T / Christianson DW | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Biochemistry / Year: 2022 Journal: Biochemistry / Year: 2022Title: Transient Prenyltransferase-Cyclase Association in Fusicoccadiene Synthase, an Assembly-Line Terpene Synthase. Authors: Jacque L Faylo / Trevor van Eeuwen / Kushol Gupta / Kenji Murakami / David W Christianson /  Abstract: Fusicoccadiene synthase from the fungus (PaFS) is an assembly-line terpene synthase that catalyzes the first two steps in the biosynthesis of Fusiccocin A, a diterpene glycoside. The C-terminal ...Fusicoccadiene synthase from the fungus (PaFS) is an assembly-line terpene synthase that catalyzes the first two steps in the biosynthesis of Fusiccocin A, a diterpene glycoside. The C-terminal prenyltransferase domain of PaFS catalyzes the condensation of one molecule of C dimethylallyl diphosphate and three molecules of C isopentenyl diphosphate to form C geranylgeranyl diphosphate, which then transits to the cyclase domain for cyclization to form fusicoccadiene. Previous structural studies of PaFS using electron microscopy (EM) revealed a central octameric prenyltransferase core with eight cyclase domains tethered in random distal positions through flexible 70-residue linkers. However, proximal prenyltransferase-cyclase configurations could be captured by covalent cross-linking and observed by cryo-EM and mass spectrometry. Here, we use cryo-EM to show that proximally configured prenyltransferase-cyclase complexes are observable even in the absence of covalent cross-linking; moreover, such complexes can involve multiple cyclase domains. A conserved basic patch on the prenyltransferase domain comprises the primary touchpoint with the cyclase domain. These results support a model for transient prenyltransferase-cyclase association in which the cyclase domains of PaFS are in facile equilibrium between proximal associated and random distal positions relative to the central prenyltransferase octamer. The results of biophysical measurements using small-angle X-ray scattering, analytical ultracentrifugation, dynamic light scattering, and size-exclusion chromatography in-line with multi-angle light scattering are consistent with this model. This model accordingly provides a framework for understanding substrate transit between the prenyltransferase and cyclase domains as well as the cooperativity observed for geranylgeranyl diphosphate cyclization. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28245.map.gz emd_28245.map.gz | 20.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28245-v30.xml emd-28245-v30.xml emd-28245.xml emd-28245.xml | 21 KB 21 KB | Display Display |  EMDB header EMDB header |
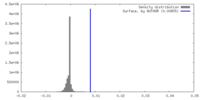
| FSC (resolution estimation) |  emd_28245_fsc.xml emd_28245_fsc.xml | 6.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_28245.png emd_28245.png | 52 KB | ||
| Masks |  emd_28245_msk_1.map emd_28245_msk_1.map | 27 MB |  Mask map Mask map | |
| Others |  emd_28245_half_map_1.map.gz emd_28245_half_map_1.map.gz emd_28245_half_map_2.map.gz emd_28245_half_map_2.map.gz | 20.7 MB 20.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28245 http://ftp.pdbj.org/pub/emdb/structures/EMD-28245 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28245 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28245 | HTTPS FTP |
-Validation report
| Summary document |  emd_28245_validation.pdf.gz emd_28245_validation.pdf.gz | 731.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_28245_full_validation.pdf.gz emd_28245_full_validation.pdf.gz | 730.7 KB | Display | |
| Data in XML |  emd_28245_validation.xml.gz emd_28245_validation.xml.gz | 12.6 KB | Display | |
| Data in CIF |  emd_28245_validation.cif.gz emd_28245_validation.cif.gz | 17.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28245 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28245 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28245 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28245 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_28245.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28245.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | PaFS prenyltransferase core with two interacting cyclase domains - East and North | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.66 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_28245_msk_1.map emd_28245_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
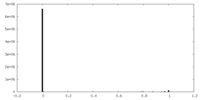
| Density Histograms |
-Half map: Half Map 1
| File | emd_28245_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
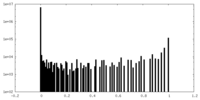
| Density Histograms |
-Half map: Half Map 2
| File | emd_28245_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
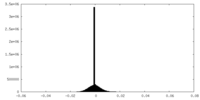
| Density Histograms |
- Sample components
Sample components
-Entire : Octameric full-length fusicoccadiene synthase prepared with metal...
| Entire | Name: Octameric full-length fusicoccadiene synthase prepared with metal cofactor and inhibitor pamidronate |
|---|---|
| Components |
|
-Supramolecule #1: Octameric full-length fusicoccadiene synthase prepared with metal...
| Supramolecule | Name: Octameric full-length fusicoccadiene synthase prepared with metal cofactor and inhibitor pamidronate type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Diaporthe amygdali (fungus) Diaporthe amygdali (fungus) |
| Recombinant expression | Organism:  |
-Macromolecule #1: Fusicoccadiene synthase
| Macromolecule | Name: Fusicoccadiene synthase / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO / EC number: fusicocca-2,10(14)-diene synthase |
|---|---|
| Source (natural) | Organism:  Diaporthe amygdali (fungus) Diaporthe amygdali (fungus) |
| Sequence | String: MGSSHHHHHH SSGLVPRGSH MEFKYSEVVE PSTYYTEGLC EGIDVRKSKF TTLEDRGAIR AHEDWNKHIG PCREYRGTLG PRFSFISVAV PECIPERLEV ISYANEFAFL HDDVTDHVGH DTGEVENDEM MTVFLEAAHT GAIDTSNKVD IRRAGKKRIQ SQLFLEMLAI ...String: MGSSHHHHHH SSGLVPRGSH MEFKYSEVVE PSTYYTEGLC EGIDVRKSKF TTLEDRGAIR AHEDWNKHIG PCREYRGTLG PRFSFISVAV PECIPERLEV ISYANEFAFL HDDVTDHVGH DTGEVENDEM MTVFLEAAHT GAIDTSNKVD IRRAGKKRIQ SQLFLEMLAI DPECAKTTMK SWARFVEVGS SRQHETRFVE LAKYIPYRIM DVGEMFWFGL VTFGLGLHIP DHELELCREL MANAWIAVGL QNDIWSWPKE RDAATLHGKD HVVNAIWVLM QEHQTDVDGA MQICRKLIVE YVAKYLEVIE ATKNDESISL DLRKYLDAML YSISGNVVWS LECPRYNPDV SFNKTQLEWM RQGLPSLESC PVLARSPEID SDESAVSPTA DESDSTEDSL GSGSRQDSSL STGLSLSPVH SNEGKDLQRV DTDHIFFEKA VLEAPYDYIA SMPSKGVRDQ FIDALNDWLR VPDVKVGKIK DAVRVLHNSS LLLDDFQDNS PLRRGKPSTH NIFGSAQTVN TATYSIIKAI GQIMEFSAGE SVQEVMNSIM ILFQGQAMDL FWTYNGHVPS EEEYYRMIDQ KTGQLFSIAT SLLLNAADNE IPRTKIQSCL HRLTRLLGRC FQICDDYQNL VSADYTKQKG FCEDLDEGKW SLALIHMIHK QRSHMALLNV LSTGRKHGGM TLEQKQFVLD IIEEEKSLDY TRSVMMDLHV QLRAEIGRIE ILLDSPNPAM RLLLELLRV |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 52.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)