[English] 日本語
 Yorodumi
Yorodumi- EMDB-2814: Electron cryotomography of the FtsZ ring in Caulobacter crescentus. -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2814 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Electron cryotomography of the FtsZ ring in Caulobacter crescentus. | |||||||||
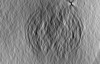
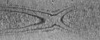
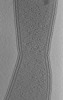
 Map data Map data | Tomogram of dividing Caulobacter crescentus CB15N grown in M2G medium. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | FtsZ / divisome / bacterial cell division / cytokinesis | |||||||||
| Biological species |  Caulobacter vibrioides (bacteria) Caulobacter vibrioides (bacteria) | |||||||||
| Method | electron tomography / cryo EM | |||||||||
 Authors Authors | Szwedziak P / Wang Q / Bharat TAM / Tsim M / Lowe J | |||||||||
 Citation Citation |  Journal: Elife / Year: 2014 Journal: Elife / Year: 2014Title: Architecture of the ring formed by the tubulin homologue FtsZ in bacterial cell division. Authors: Piotr Szwedziak / Qing Wang / Tanmay A M Bharat / Matthew Tsim / Jan Löwe /  Abstract: Membrane constriction is a prerequisite for cell division. The most common membrane constriction system in prokaryotes is based on the tubulin homologue FtsZ, whose filaments in E. coli are anchored ...Membrane constriction is a prerequisite for cell division. The most common membrane constriction system in prokaryotes is based on the tubulin homologue FtsZ, whose filaments in E. coli are anchored to the membrane by FtsA and enable the formation of the Z-ring and divisome. The precise architecture of the FtsZ ring has remained enigmatic. In this study, we report three-dimensional arrangements of FtsZ and FtsA filaments in C. crescentus and E. coli cells and inside constricting liposomes by means of electron cryomicroscopy and cryotomography. In vivo and in vitro, the Z-ring is composed of a small, single-layered band of filaments parallel to the membrane, creating a continuous ring through lateral filament contacts. Visualisation of the in vitro reconstituted constrictions as well as a complete tracing of the helical paths of the filaments with a molecular model favour a mechanism of FtsZ-based membrane constriction that is likely to be accompanied by filament sliding. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2814.map.gz emd_2814.map.gz | 348.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2814-v30.xml emd-2814-v30.xml emd-2814.xml emd-2814.xml | 8.9 KB 8.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd-2814.png emd-2814.png | 521.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2814 http://ftp.pdbj.org/pub/emdb/structures/EMD-2814 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2814 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2814 | HTTPS FTP |
-Validation report
| Summary document |  emd_2814_validation.pdf.gz emd_2814_validation.pdf.gz | 170.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2814_full_validation.pdf.gz emd_2814_full_validation.pdf.gz | 169.4 KB | Display | |
| Data in XML |  emd_2814_validation.xml.gz emd_2814_validation.xml.gz | 4.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2814 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2814 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2814 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2814 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2814.map.gz / Format: CCP4 / Size: 370.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2814.map.gz / Format: CCP4 / Size: 370.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Tomogram of dividing Caulobacter crescentus CB15N grown in M2G medium. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 17.98 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
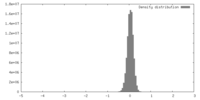
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Caulobacter crescentus CB15N grown in M2G medium.
| Entire | Name: Caulobacter crescentus CB15N grown in M2G medium. |
|---|---|
| Components |
|
-Supramolecule #1000: Caulobacter crescentus CB15N grown in M2G medium.
| Supramolecule | Name: Caulobacter crescentus CB15N grown in M2G medium. / type: sample / ID: 1000 Details: Dividing Caulobacter crescentus CB15N grown in M2G medium. Number unique components: 1 |
|---|
-Supramolecule #1: Caulobacter crescentus
| Supramolecule | Name: Caulobacter crescentus / type: organelle_or_cellular_component / ID: 1 Details: Cells were grown to mid-log phase, mixed with 10 nm protein A gold. Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  Caulobacter vibrioides (bacteria) / Strain: CB15N / Cell: Bacterial Caulobacter vibrioides (bacteria) / Strain: CB15N / Cell: Bacterial |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | Details: M2G medium |
|---|---|
| Grid | Details: Quantifoil R3.5/1 300 mesh Cu/Rh holey carbon grid |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV / Method: 2.5 s blot on Whatmann 1 filter paper. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Alignment procedure | Legacy - Astigmatism: Visualisation of power spectrum. |
| Specialist optics | Energy filter - Name: Gatan Quantum / Energy filter - Lower energy threshold: 0.0 eV / Energy filter - Upper energy threshold: 20.0 eV |
| Date | Jul 31, 2014 |
| Image recording | Category: CCD / Film or detector model: GATAN K2 (4k x 4k) / Number real images: 121 / Average electron dose: 150 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 26000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: -10.0 µm / Nominal defocus min: -10.0 µm / Nominal magnification: 26000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Tilt series - Axis1 - Min angle: 0 ° / Tilt series - Axis1 - Max angle: 60 ° / Tilt series - Axis1 - Angle increment: 1 ° |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | Tilt series data was aligned using IMOD and reconstruction was carried out using tomo3D. |
|---|---|
| Final reconstruction | Algorithm: OTHER / Software - Name: IMOD, tomo3D / Number images used: 121 |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)