+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Cryo-EM structure of consensus elemental paused elongation complex with an unfolded TL | |||||||||
 マップデータ マップデータ | a postprocessed map | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | RNA polymerase / cryo-EM structure / elemental pause / Escherichia coli / transcription / TRANSFERASE-DNA-RNA complex / transcriptional regulation / transcriptional pausing | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報DNA-directed RNA polymerase complex / ribonucleoside binding / DNA-directed RNA polymerase / DNA-directed RNA polymerase activity / protein dimerization activity / DNA-templated transcription / magnesium ion binding / DNA binding / zinc ion binding / cytoplasm / cytosol 類似検索 - 分子機能 | |||||||||
| 生物種 |  | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.8 Å | |||||||||
 データ登録者 データ登録者 | Kang JY / Chen J / Llewellyn E / Landick R / Darst SA | |||||||||
| 資金援助 |  米国, 2件 米国, 2件
| |||||||||
 引用 引用 |  ジャーナル: Proc Natl Acad Sci U S A / 年: 2023 ジャーナル: Proc Natl Acad Sci U S A / 年: 2023タイトル: An ensemble of interconverting conformations of the elemental paused transcription complex creates regulatory options. 著者: Jin Young Kang / Tatiana V Mishanina / Yu Bao / James Chen / Eliza Llewellyn / James Liu / Seth A Darst / Robert Landick /   要旨: Transcriptional pausing underpins the regulation of cellular RNA synthesis, but its mechanism remains incompletely understood. Sequence-specific interactions of DNA and RNA with the dynamic, ...Transcriptional pausing underpins the regulation of cellular RNA synthesis, but its mechanism remains incompletely understood. Sequence-specific interactions of DNA and RNA with the dynamic, multidomain RNA polymerase (RNAP) trigger reversible conformational changes at pause sites that temporarily interrupt the nucleotide addition cycle. These interactions initially rearrange the elongation complex (EC) into an elemental paused EC (ePEC). ePECs can form longer-lived PECs by further rearrangements or interactions of diffusible regulators. For both bacterial and mammalian RNAPs, a half-translocated state in which the next DNA template base fails to load into the active site appears central to the ePEC. Some RNAPs also swivel interconnected modules that may stabilize the ePEC. However, it is unclear whether swiveling and half-translocation are requisite features of a single ePEC state or if multiple ePEC states exist. Here, we use cryo-electron microscopy (cryo-EM) analysis of ePECs with different RNA-DNA sequences combined with biochemical probes of ePEC structure to define an interconverting ensemble of ePEC states. ePECs occupy either pre- or half-translocated states but do not always swivel, indicating that difficulty in forming the posttranslocated state at certain RNA-DNA sequences may be the essence of the ePEC. The existence of multiple ePEC conformations has broad implications for transcriptional regulation. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_28113.map.gz emd_28113.map.gz | 60 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-28113-v30.xml emd-28113-v30.xml emd-28113.xml emd-28113.xml | 32.8 KB 32.8 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_28113.png emd_28113.png | 151.9 KB | ||
| Filedesc metadata |  emd-28113.cif.gz emd-28113.cif.gz | 9.4 KB | ||
| その他 |  emd_28113_additional_1.map.gz emd_28113_additional_1.map.gz emd_28113_half_map_1.map.gz emd_28113_half_map_1.map.gz emd_28113_half_map_2.map.gz emd_28113_half_map_2.map.gz | 51.4 MB 49.7 MB 49.7 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28113 http://ftp.pdbj.org/pub/emdb/structures/EMD-28113 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28113 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28113 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  8egbMC  8eg7C  8eg8C  8eh8C  8eh9C  8ehaC  8ehfC  8ehiC C: 同じ文献を引用 ( M: このマップから作成された原子モデル |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_28113.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_28113.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | a postprocessed map | ||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.3 Å | ||||||||||||||||||||||||||||||||||||
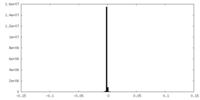
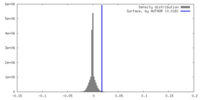
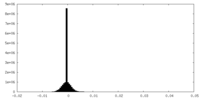
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-追加マップ: a local-filtered map
| ファイル | emd_28113_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | a local-filtered map | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
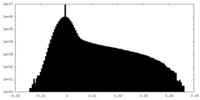
| 密度ヒストグラム |
-ハーフマップ: a second half map
| ファイル | emd_28113_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | a second half map | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: a first half mpa
| ファイル | emd_28113_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | a first half mpa | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
+全体 : consensus elemental paused elongation complex
+超分子 #1: consensus elemental paused elongation complex
+分子 #1: non-template DNA
+分子 #2: template DNA
+分子 #3: RNA
+分子 #4: DNA-directed RNA polymerase subunit alpha
+分子 #5: DNA-directed RNA polymerase subunit beta
+分子 #6: DNA-directed RNA polymerase subunit beta'
+分子 #7: DNA-directed RNA polymerase subunit omega
+分子 #8: CHAPSO
+分子 #9: MAGNESIUM ION
+分子 #10: ZINC ION
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 8 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 緩衝液 | pH: 8 構成要素:
| |||||||||||||||
| グリッド | モデル: C-flat-1.2/1.3 / 材質: GOLD / メッシュ: 400 / 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: HOLEY / 前処理 - タイプ: GLOW DISCHARGE / 前処理 - 時間: 20 sec. / 前処理 - 雰囲気: AIR | |||||||||||||||
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 22 K / 装置: FEI VITROBOT MARK IV / 詳細: blot force 0, blot time 3 s. |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 検出モード: SUPER-RESOLUTION / デジタル化 - サイズ - 横: 3710 pixel / デジタル化 - サイズ - 縦: 3838 pixel / デジタル化 - 画像ごとのフレーム数: 1-50 / 撮影したグリッド数: 1 / 実像数: 2525 / 平均露光時間: 10.0 sec. / 平均電子線量: 47.34 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / Cs: 2.7 mm / 最大 デフォーカス(公称値): 2.4 µm / 最小 デフォーカス(公称値): 0.8 µm |
| 試料ステージ | 試料ホルダーモデル: FEI TITAN KRIOS AUTOGRID HOLDER |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
+ 画像解析
画像解析
-原子モデル構築 1
| 初期モデル |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 精密化 | 空間: REAL / プロトコル: RIGID BODY FIT | ||||||||||||
| 得られたモデル |  PDB-8egb: |
 ムービー
ムービー コントローラー
コントローラー













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)