[English] 日本語
 Yorodumi
Yorodumi- EMDB-28065: Cas7-11 in complex with DR-mismatched target RNA, Csx29 and Csx30 -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cas7-11 in complex with DR-mismatched target RNA, Csx29 and Csx30 | |||||||||
 Map data Map data | Cas7-11 in complex with DR-mismatched target RNA, Csx29 and Csx30, focus refined on Cas7-11 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CRISPR / endonuclease / endopeptidase / RNA BINDING PROTEIN-RNA complex | |||||||||
| Function / homology | CHAT domain / CHAT domain / : / CRISPR type III-associated protein / RAMP superfamily / defense response to virus / CRISPR-associated RAMP family protein / Uncharacterized protein / CHAT domain-containing protein Function and homology information Function and homology information | |||||||||
| Biological species |  Desulfonema ishimotonii (bacteria) / synthetic construct (others) Desulfonema ishimotonii (bacteria) / synthetic construct (others) | |||||||||
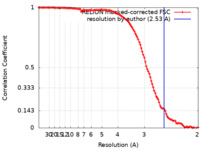
| Method | single particle reconstruction / cryo EM / Resolution: 2.53 Å | |||||||||
 Authors Authors | Demircioglu FE / Wilkinson ME / Strecker J / Li D / Faure G / Macrae RK / Zhang F | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2022 Journal: Science / Year: 2022Title: RNA-activated protein cleavage with a CRISPR-associated endopeptidase. Authors: Jonathan Strecker / F Esra Demircioglu / David Li / Guilhem Faure / Max E Wilkinson / Jonathan S Gootenberg / Omar O Abudayyeh / Hiroshi Nishimasu / Rhiannon K Macrae / Feng Zhang /   Abstract: In prokaryotes, CRISPR-Cas systems provide adaptive immune responses against foreign genetic elements through RNA-guided nuclease activity. Recently, additional genes with non-nuclease functions have ...In prokaryotes, CRISPR-Cas systems provide adaptive immune responses against foreign genetic elements through RNA-guided nuclease activity. Recently, additional genes with non-nuclease functions have been found in genetic association with CRISPR systems, suggesting that there may be other RNA-guided non-nucleolytic enzymes. One such gene from encodes the TPR-CHAT protease Csx29, which is associated with the CRISPR effector Cas7-11. Here, we demonstrate that this CRISPR-associated protease (CASP) exhibits programmable RNA-activated endopeptidase activity against a sigma factor inhibitor to regulate a transcriptional response. Cryo-electron microscopy of an active and substrate-bound CASP complex reveals an allosteric activation mechanism that reorganizes Csx29 catalytic residues upon target RNA binding. This work reveals an RNA-guided function in nature that can be leveraged for RNA-sensing applications in vitro and in human cells. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28065.map.gz emd_28065.map.gz | 116.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28065-v30.xml emd-28065-v30.xml emd-28065.xml emd-28065.xml | 20.9 KB 20.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_28065_fsc.xml emd_28065_fsc.xml | 11.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_28065.png emd_28065.png | 605.2 KB | ||
| Masks |  emd_28065_msk_1.map emd_28065_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-28065.cif.gz emd-28065.cif.gz | 7.8 KB | ||
| Others |  emd_28065_half_map_1.map.gz emd_28065_half_map_1.map.gz emd_28065_half_map_2.map.gz emd_28065_half_map_2.map.gz | 98.3 MB 98.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28065 http://ftp.pdbj.org/pub/emdb/structures/EMD-28065 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28065 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28065 | HTTPS FTP |
-Validation report
| Summary document |  emd_28065_validation.pdf.gz emd_28065_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_28065_full_validation.pdf.gz emd_28065_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_28065_validation.xml.gz emd_28065_validation.xml.gz | 18.5 KB | Display | |
| Data in CIF |  emd_28065_validation.cif.gz emd_28065_validation.cif.gz | 24.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28065 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28065 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28065 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28065 | HTTPS FTP |
-Related structure data
| Related structure data |  8eeyMC  8eexC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_28065.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28065.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cas7-11 in complex with DR-mismatched target RNA, Csx29 and Csx30, focus refined on Cas7-11 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.9945 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_28065_msk_1.map emd_28065_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Cas7-11 in complex with DR-mismatched target RNA, Csx29...
| File | emd_28065_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cas7-11 in complex with DR-mismatched target RNA, Csx29 and Csx30, half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Cas7-11 in complex with DR-mismatched target RNA, Csx29...
| File | emd_28065_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cas7-11 in complex with DR-mismatched target RNA, Csx29 and Csx30, half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cas7-11 in complex with DR-mismatched target RNA, Csx29 and Csx30
| Entire | Name: Cas7-11 in complex with DR-mismatched target RNA, Csx29 and Csx30 |
|---|---|
| Components |
|
-Supramolecule #1: Cas7-11 in complex with DR-mismatched target RNA, Csx29 and Csx30
| Supramolecule | Name: Cas7-11 in complex with DR-mismatched target RNA, Csx29 and Csx30 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Desulfonema ishimotonii (bacteria) Desulfonema ishimotonii (bacteria) |
-Macromolecule #1: Csx30
| Macromolecule | Name: Csx30 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Desulfonema ishimotonii (bacteria) Desulfonema ishimotonii (bacteria) |
| Molecular weight | Theoretical: 62.688184 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MNTTTYNTTT DALLEWGKVY FQKEDFSEFL DNLEAYISDA GDSLKDELES GVEKLVLGIK SAEAVIFGEA VIGTTPENEA WYDAEESFL TLDCAVWLSQ ALDRVVRRQD ASLADSLIAR LDEAINRVAE KLYADNLSPL RFSSLNEIRR SALEATDEKY H YLFPWHGA ...String: MNTTTYNTTT DALLEWGKVY FQKEDFSEFL DNLEAYISDA GDSLKDELES GVEKLVLGIK SAEAVIFGEA VIGTTPENEA WYDAEESFL TLDCAVWLSQ ALDRVVRRQD ASLADSLIAR LDEAINRVAE KLYADNLSPL RFSSLNEIRR SALEATDEKY H YLFPWHGA ACDVDENILL ILTEEYHLIG ADKAGANLSE ELRGDLPFIF AELERDEVLR AYVEKENALS LALENTMREH WA FGLLEAA RDEGYNHPYP ADVGMRIHQV ARAVFSQTNL SPAERLAVAI AGACFTPEIS EDRRLEILLD CEERVCEIEA PTG DDTSVR VIKDLKALAD HRVRHEIPAE SLVSLWFEQI EAAGTDFDTK TPMDELVLRM LSDNVITLSV DRKAASQTET DDVK PQKGK IIPFPVPDIA NDEVEYQKAV GGGSANDSKV KFPGLLEIQG CRDGDKAILL EDTDDAAANH RKLFSILKAG KLNSA FFIQ SDDGEWVESE SKPTMEDNRI ILHDSHHSSF VWILDTGSMQ LRQSVKCVKD ALNKKTGSAK KLKPKTMIVW VTIPQE G UniProtKB: Uncharacterized protein, Uncharacterized protein |
-Macromolecule #2: Cas7-11
| Macromolecule | Name: Cas7-11 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Desulfonema ishimotonii (bacteria) Desulfonema ishimotonii (bacteria) |
| Molecular weight | Theoretical: 183.844984 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MTTTMKISIE FLEPFRMTKW QESTRRNKNN KEFVRGQAFA RWHRNKKDNT KGRPYITGTL LRSAVIRSAE NLLTLSDGKI SEKTCCPGK FDTEDKDRLL QLRQRSTLRW TDKNPCPDNA ETYCPFCELL GRSGNDGKKA EKKDWRFRIH FGNLSLPGKP D FDGPKAIG ...String: MTTTMKISIE FLEPFRMTKW QESTRRNKNN KEFVRGQAFA RWHRNKKDNT KGRPYITGTL LRSAVIRSAE NLLTLSDGKI SEKTCCPGK FDTEDKDRLL QLRQRSTLRW TDKNPCPDNA ETYCPFCELL GRSGNDGKKA EKKDWRFRIH FGNLSLPGKP D FDGPKAIG SQRVLNRVDF KSGKAHDFFK AYEVDHTRFP RFEGEITIDN KVSAEARKLL CDSLKFTDRL CGALCVIRFD EY TPAADSG KQTENVQAEP NANLAEKTAE QIISILDDNK KTEYTRLLAD AIRSLRRSSK LVAGLPKDHD GKDDHYLWDI GKK KKDENS VTIRQILTTS ADTKELKNAG KWREFCEKLG EALYLKSKDM SGGLKITRRI LGDAEFHGKP DRLEKSRSVS IGSV LKETV VCGELVAKTP FFFGAIDEDA KQTALQVLLT PDNKYRLPRS AVRGILRRDL QTYFDSPCNA ELGGRPCMCK TCRIM RGIT VMDARSEYNA PPEIRHRTRI NPFTGTVAEG ALFNMEVAPE GIVFPFQLRY RGSEDGLPDA LKTVLKWWAE GQAFMS GAA STGKGRFRME NAKYETLDLS DENQRNDYLK NWGWRDEKGL EELKKRLNSG LPEPGNYRDP KWHEINVSIE MASPFIN GD PIRAAVDKRG TAVVTFVKYK AEGEEAKPVC AYKAESFRGV IRSAVARIHM EDGVPLTELT HSDCECLLCQ IFGSEYEA G KIRFEDLVFE SDPEPVTFDH VAIDRFTGGA ADKKKFDDSP LPGSPARPLM LKGSFWIRRD VLEDEEYCKA LGKALADVN NGLYPLGGKS AIGYGQVKSL GIKGDDKRIS RLMNPAFDET DVAVPEKPKT DAEVRIEAEK VYYPHYFVEP HKKVEREEKP CGHQKFHEG RLTGKIRCKL ITKTPLIVPD TSNDDFFRPA DKEARKEKDE YHKSYAFFRL HKQIMIPGSE LRGMVSSVYE T VTNSCFRI FDETKRLSWR MDADHQNVLQ DFLPGRVTAD GKHIQKFSET ARVPFYDKTQ KHFDILDEQE IAGEKPVRMW VK RFIKRLS LVDPAKHPQK KQDNKWKRRK EGIATFIEQK NGSYYFNVVT NNGCTSFHLW HKPDNFDQEK LEGIQNGEKL DCW VRDSRY QKAFQEIPEN DPDGWECKEG YLHVVGPSKV EFSDKKGDVI NNFQGTLPSV PNDWKTIRTN DFKNRKRKNE PVFC CEDDK GNYYTMAKYC ETFFFDLKEN EEYEIPEKAR IKYKELLRVY NNNPQAVPES VFQSRVAREN VEKLKSGDLV YFKHN EKYV EDIVPVRISR TVDDRMIGKR MSADLRPCHG DWVEDGDLSA LNAYPEKRLL LRHPKGLCPA CRLFGTGSYK GRVRFG FAS LENDPEWLIP GKNPGDPFHG GPVMLSLLER PRPTWSIPGS DNKFKVPGRK FYVHHHAWKT IKDGNHPTTG KAIEQSP NN RTVEALAGGN SFSFEIAFEN LKEWELGLLI HSLQLEKGLA HKLGMAKSMG FGSVEIDVES VRLRKDWKQW RNGNSEIP N WLGKGFAKLK EWFRDELDFI ENLKKLLWFP EGDQAPRVCY PMLRKKDDPN GNSGYEELKD GEFKKEDRQK KLTTPWTPW A UniProtKB: CRISPR-associated RAMP family protein |
-Macromolecule #3: Csx29
| Macromolecule | Name: Csx29 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Desulfonema ishimotonii (bacteria) Desulfonema ishimotonii (bacteria) |
| Molecular weight | Theoretical: 88.311875 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSNPIRDIQD RLKTAKFDNK DDMMNLASSL YKYEKQLMDS SEATLCQQGL SNRPNSFSQL SQFRDSDIQS KAGGQTGKFW QNEYEACKN FQTHKERRET LEQIIRFLQN GAEEKDADDL LLKTLARAYF HRGLLYRPKG FSVPARKVEA MKKAIAYCEI I LDKNEEES ...String: MSNPIRDIQD RLKTAKFDNK DDMMNLASSL YKYEKQLMDS SEATLCQQGL SNRPNSFSQL SQFRDSDIQS KAGGQTGKFW QNEYEACKN FQTHKERRET LEQIIRFLQN GAEEKDADDL LLKTLARAYF HRGLLYRPKG FSVPARKVEA MKKAIAYCEI I LDKNEEES EALRIWLYAA MELRRCGEEY PENFAEKLFY LANDGFISEL YDIRLFLEYT EREEDNNFLD MILQENQDRE RL FELCLYK ARACFHLNQL NDVRIYGESA IDNAPGAFAD PFWDELVEFI RMLRNKKSEL WKEIAIKAWD KCREKEMKVG NNI YLSWYW ARQRELYDLA FMAQDGIEKK TRIADSLKSR TTLRIQELNE LRKDAHRKQN RRLEDKLDRI IEQENEARDG AYLR RNPPC FTGGKREEIP FARLPQNWIA VHFYLNELES HEGGKGGHAL IYDPQKAEKD QWQDKSFDYK ELHRKFLEWQ ENYIL NEEG SADFLVTLCR EIEKAMPFLF KSEVIPEDRP VLWIPHGFLH RLPLHAAMKS GNNSNIEIFW ERHASRYLPA WHLFDP APY SREESSTLLK NFEEYDFQNL ENGEIEVYAP SSPKKVKEAI RENPAILLLL CHGEADMTNP FRSCLKLKNK DMTIFDL LT VEDVRLSGSR ILLGACESDM VPPLEFSVDE HLSVSGAFLS HKAGEIVAGL WTVDSEKVDE CYSYLVEEKD FLRNLQEW Q MAETENFRSE NDSSLFYKIA PFRIIGFPAE UniProtKB: CHAT domain-containing protein |
-Macromolecule #4: crRNA
| Macromolecule | Name: crRNA / type: rna / ID: 4 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Desulfonema ishimotonii (bacteria) Desulfonema ishimotonii (bacteria) |
| Molecular weight | Theoretical: 12.134133 KDa |
| Sequence | String: UUGAUGUCAC GGAACCUUUG UUGUCUUCGA CAUGGGUA |
-Macromolecule #5: DR-mismatched target RNA
| Macromolecule | Name: DR-mismatched target RNA / type: rna / ID: 5 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 8.931381 KDa |
| Sequence | String: UACCCAUGUC GAAGACAACA AAGUAUUU |
-Macromolecule #6: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 6 / Number of copies: 4 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)