+ Open data
Open data
- Basic information
Basic information
| Entry | 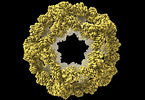 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
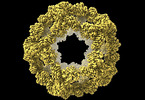
| Title | Structure of a nanoparticle with icosahedral symmetry | |||||||||
 Map data Map data | Post-processed map | |||||||||
 Sample Sample |
| |||||||||
| Function / homology | 4-hydroxy-2-oxoglutarate aldolase / 2-dehydro-3-deoxy-phosphogluconate aldolase / 2-dehydro-3-deoxy-phosphogluconate aldolase activity / KDPG/KHG aldolase / KDPG and KHG aldolase / (R,S)-4-hydroxy-2-oxoglutarate aldolase activity / Aldolase-type TIM barrel / 4-Hydroxy-2-oxoglutarate aldolase / 2-dehydro-3-deoxyphosphogluconate aldolase Function and homology information Function and homology information | |||||||||
| Biological species |   Thermotoga maritima MSB8 (bacteria) Thermotoga maritima MSB8 (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | McCarthy S / Gonen S | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Heliyon / Year: 2022 Journal: Heliyon / Year: 2022Title: Improved interface packing and design opportunities revealed by CryoEM analysis of a designed protein nanocage. Authors: Stephen McCarthy / Shane Gonen /  Abstract: Symmetric protein assemblies play important roles in nature which makes them an attractive target for engineering. symmetric protein complexes can be created through computational protein design to ...Symmetric protein assemblies play important roles in nature which makes them an attractive target for engineering. symmetric protein complexes can be created through computational protein design to tailor their properties from first principles, and recently several protein nanocages have been created by bringing together protein components through hydrophobic interactions. Accurate experimental structures of newly-developed proteins are essential to validate their design, improve assembly stability, and tailor downstream applications. We describe the CryoEM structure of the nanocage I3-01, at an overall resolution of 3.5 Å. I3-01, comprising 60 aldolase subunits arranged with icosahedral symmetry, has resisted high-resolution characterization. Some key differences between the refined structure and the original design are identified, such as improved packing of hydrophobic sidechains, providing insight to the resistance of I3-01 to high-resolution averaging. Based on our analysis, we suggest factors important in the design and structural processing of new assemblies. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28027.map.gz emd_28027.map.gz | 17.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28027-v30.xml emd-28027-v30.xml emd-28027.xml emd-28027.xml | 13.4 KB 13.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_28027.png emd_28027.png | 128.3 KB | ||
| Others |  emd_28027_half_map_1.map.gz emd_28027_half_map_1.map.gz emd_28027_half_map_2.map.gz emd_28027_half_map_2.map.gz | 140.9 MB 140.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28027 http://ftp.pdbj.org/pub/emdb/structures/EMD-28027 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28027 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28027 | HTTPS FTP |
-Validation report
| Summary document |  emd_28027_validation.pdf.gz emd_28027_validation.pdf.gz | 605.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_28027_full_validation.pdf.gz emd_28027_full_validation.pdf.gz | 604.9 KB | Display | |
| Data in XML |  emd_28027_validation.xml.gz emd_28027_validation.xml.gz | 14.9 KB | Display | |
| Data in CIF |  emd_28027_validation.cif.gz emd_28027_validation.cif.gz | 17.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28027 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28027 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28027 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28027 | HTTPS FTP |
-Related structure data
| Related structure data |  8ed3MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_28027.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28027.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Post-processed map | ||||||||||||||||||||
| Voxel size | X=Y=Z: 1.31 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half-map 1
| File | emd_28027_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half-map 2
| File | emd_28027_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : I3-01
| Entire | Name: I3-01 |
|---|---|
| Components |
|
-Supramolecule #1: I3-01
| Supramolecule | Name: I3-01 / type: complex / ID: 1 / Chimera: Yes / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Thermotoga maritima MSB8 (bacteria) Thermotoga maritima MSB8 (bacteria) |
-Macromolecule #1: Designed I3-01 icosahedron
| Macromolecule | Name: Designed I3-01 icosahedron / type: protein_or_peptide / ID: 1 / Number of copies: 60 / Enantiomer: LEVO / EC number: 2-dehydro-3-deoxy-phosphogluconate aldolase |
|---|---|
| Source (natural) | Organism:   Thermotoga maritima MSB8 (bacteria) Thermotoga maritima MSB8 (bacteria)Strain: ATCC 43589 / DSM 3109 / JCM 10099 / NBRC 100826 / MSB8 |
| Molecular weight | Theoretical: 21.660584 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MEELFKKHKI VAVLRANSVE EAKKKALAVF LGGVHLIEIT FTVPDADTVI KELSFLKEMG AIIGAGTVTS VEQCRKAVES GAEFIVSPH LDEEISQFCK EKGVFYMPGV MTPTELVKAM KLGHTILKLF PGEVVGPQFV KAMKGPFPNV KFVPTGGVNL D NVCEWFKA ...String: MEELFKKHKI VAVLRANSVE EAKKKALAVF LGGVHLIEIT FTVPDADTVI KELSFLKEMG AIIGAGTVTS VEQCRKAVES GAEFIVSPH LDEEISQFCK EKGVFYMPGV MTPTELVKAM KLGHTILKLF PGEVVGPQFV KAMKGPFPNV KFVPTGGVNL D NVCEWFKA GVLAVGVGSA LVKGTPVEVA EKAKAFVEKI RGC |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 2.1 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER / Details: Stochastic gradient descent from 2D averages |
|---|---|
| Final reconstruction | Applied symmetry - Point group: I (icosahedral) / Resolution.type: BY AUTHOR / Resolution: 3.5 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 147349 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-8ed3: |
 Movie
Movie Controller
Controller





 Z
Z Y
Y X
X