[English] 日本語
 Yorodumi
Yorodumi- EMDB-27745: Cryo-EM structure of RIG-I bound to the internal sites of OHSLR30... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of RIG-I bound to the internal sites of OHSLR30 (+ATP) | |||||||||
 Map data Map data | Cryo-EM structure of protein-RNA complex 9 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ribonucleoprotein complex / RNA sensor / RIG-I like receptor / Hydrolase-RNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of type III interferon production / RIG-I signaling pathway / positive regulation of myeloid dendritic cell cytokine production / OAS antiviral response / detection of virus / NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 / positive regulation of response to cytokine stimulus / positive regulation of granulocyte macrophage colony-stimulating factor production / pattern recognition receptor activity / TRAF6 mediated IRF7 activation ...regulation of type III interferon production / RIG-I signaling pathway / positive regulation of myeloid dendritic cell cytokine production / OAS antiviral response / detection of virus / NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 / positive regulation of response to cytokine stimulus / positive regulation of granulocyte macrophage colony-stimulating factor production / pattern recognition receptor activity / TRAF6 mediated IRF7 activation / cytoplasmic pattern recognition receptor signaling pathway / cellular response to exogenous dsRNA / RSV-host interactions / response to exogenous dsRNA / positive regulation of interferon-alpha production / TRAF6 mediated NF-kB activation / bicellular tight junction / positive regulation of defense response to virus by host / antiviral innate immune response / positive regulation of interferon-beta production / regulation of cell migration / positive regulation of interleukin-8 production / Negative regulators of DDX58/IFIH1 signaling / DDX58/IFIH1-mediated induction of interferon-alpha/beta / response to virus / Evasion by RSV of host interferon responses / ISG15 antiviral mechanism / ruffle membrane / positive regulation of interleukin-6 production / SARS-CoV-1 activates/modulates innate immune responses / positive regulation of tumor necrosis factor production / double-stranded RNA binding / Ovarian tumor domain proteases / actin cytoskeleton / TRAF3-dependent IRF activation pathway / gene expression / double-stranded DNA binding / defense response to virus / RNA helicase activity / single-stranded RNA binding / Ub-specific processing proteases / RNA helicase / ribonucleoprotein complex / innate immune response / ubiquitin protein ligase binding / positive regulation of gene expression / GTP binding / SARS-CoV-2 activates/modulates innate and adaptive immune responses / positive regulation of transcription by RNA polymerase II / ATP hydrolysis activity / zinc ion binding / ATP binding / identical protein binding / cytosol / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
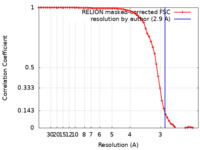
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||
 Authors Authors | Wang W / Pyle AM | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2022 Journal: Mol Cell / Year: 2022Title: The RIG-I receptor adopts two different conformations for distinguishing host from viral RNA ligands. Authors: Wenshuai Wang / Anna Marie Pyle /  Abstract: RIG-I is an essential innate immune receptor for detecting and responding to infection by RNA viruses. RIG-I specifically recognizes the unique molecular features of viral RNA molecules and ...RIG-I is an essential innate immune receptor for detecting and responding to infection by RNA viruses. RIG-I specifically recognizes the unique molecular features of viral RNA molecules and selectively distinguishes them from closely related RNAs abundant in host cells. The physical basis for this exquisite selectivity is revealed through a series of high-resolution cryo-EM structures of RIG-I in complex with host and viral RNA ligands. These studies demonstrate that RIG-I actively samples double-stranded RNAs in the cytoplasm and distinguishes them by adopting two different types of protein folds. Upon binding viral RNA, RIG-I adopts a high-affinity conformation that is conducive to signaling, while host RNA induces an autoinhibited conformation that stimulates RNA release. By coupling protein folding with RNA binding selectivity, RIG-I distinguishes RNA molecules that differ by as little as one phosphate group, thereby explaining the molecular basis for selective antiviral sensing and the induction of autoimmunity upon RIG-I dysregulation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27745.map.gz emd_27745.map.gz | 19.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27745-v30.xml emd-27745-v30.xml emd-27745.xml emd-27745.xml | 16.4 KB 16.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27745_fsc.xml emd_27745_fsc.xml | 6.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_27745.png emd_27745.png | 51.9 KB | ||
| Masks |  emd_27745_msk_1.map emd_27745_msk_1.map | 20.8 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-27745.cif.gz emd-27745.cif.gz | 6.2 KB | ||
| Others |  emd_27745_half_map_1.map.gz emd_27745_half_map_1.map.gz emd_27745_half_map_2.map.gz emd_27745_half_map_2.map.gz | 15.9 MB 15.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27745 http://ftp.pdbj.org/pub/emdb/structures/EMD-27745 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27745 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27745 | HTTPS FTP |
-Validation report
| Summary document |  emd_27745_validation.pdf.gz emd_27745_validation.pdf.gz | 766.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_27745_full_validation.pdf.gz emd_27745_full_validation.pdf.gz | 765.7 KB | Display | |
| Data in XML |  emd_27745_validation.xml.gz emd_27745_validation.xml.gz | 11.8 KB | Display | |
| Data in CIF |  emd_27745_validation.cif.gz emd_27745_validation.cif.gz | 16 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27745 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27745 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27745 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27745 | HTTPS FTP |
-Related structure data
| Related structure data |  8dvuMC  7tnxC  7tnyC  7tnzC  7to0C  7to1C  7to2C  8dvrC  8dvsC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27745.map.gz / Format: CCP4 / Size: 20.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27745.map.gz / Format: CCP4 / Size: 20.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of protein-RNA complex 9 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.149 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_27745_msk_1.map emd_27745_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map 1
| File | emd_27745_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map 2
| File | emd_27745_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of RIG-I with OHSLR30
| Entire | Name: Complex of RIG-I with OHSLR30 |
|---|---|
| Components |
|
-Supramolecule #1: Complex of RIG-I with OHSLR30
| Supramolecule | Name: Complex of RIG-I with OHSLR30 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Antiviral innate immune response receptor RIG-I
| Macromolecule | Name: Antiviral innate immune response receptor RIG-I / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: RNA helicase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 106.740555 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MTTEQRRSLQ AFQDYIRKTL DPTYILSYMA PWFREEEVQY IQAEKNNKGP MEAATLFLKF LLELQEEGWF RGFLDALDHA GYSGLYEAI ESWDFKKIEK LEEYRLLLKR LQPEFKTRII PTDIISDLSE CLINQECEEI LQICSTKGMM AGAEKLVECL L RSDKENWP ...String: MTTEQRRSLQ AFQDYIRKTL DPTYILSYMA PWFREEEVQY IQAEKNNKGP MEAATLFLKF LLELQEEGWF RGFLDALDHA GYSGLYEAI ESWDFKKIEK LEEYRLLLKR LQPEFKTRII PTDIISDLSE CLINQECEEI LQICSTKGMM AGAEKLVECL L RSDKENWP KTLKLALEKE RNKFSELWIV EKGIKDVETE DLEDKMETSD IQIFYQEDPE CQNLSENSCP PSEVSDTNLY SP FKPRNYQ LELALPAMKG KNTIICAPTG CGKTFVSLLI CEHHLKKFPQ GQKGKVVFFA NQIPVYEQQK SVFSKYFERH GYR VTGISG ATAENVPVEQ IVENNDIIIL TPQILVNNLK KGTIPSLSIF TLMIFDECHN TSKQHPYNMI MFNYLDQKLG GSSG PLPQV IGLTASVGVG DAKNTDEALD YICKLCASLD ASVIATVKHN LEELEQVVYK PQKFFRKVES RISDKFKYII AQLMR DTES LAKRICKDLE NLSQIQNREF GTQKYEQWIV TVQKACMVFQ MPDKDEESRI CKALFLYTSH LRKYNDALII SEHARM KDA LDYLKDFFSN VRAAGFDEIE QDLTQRFEEK LQELESVSRD PSNENPKLED LCFILQEEYH LNPETITILF VKTRALV DA LKNWIEGNPK LSFLKPGILT GRGKTNQNTG MTLPAQKCIL DAFKASGDHN ILIATSVADE GIDIAQCNLV ILYEYVGN V IKMIQTRGRG RARGSKCFLL TSNAGVIEKE QINMYKEKMM NDSILRLQTW DEAVFREKIL HIQTHEKFIR DSQEKPKPV PDKENKKLLC RKCKALACYT ADVRVIEECH YTVLGDAFKE CFVSRPHPKP KQFSSFEKRA KIFCARQNCS HDWGIHVKYK TFEIPVIKI ESFVVEDIAT GVQTLYSKWK DFHFEKIPFD PAEMSK UniProtKB: Antiviral innate immune response receptor RIG-I |
-Macromolecule #2: OHSLR30
| Macromolecule | Name: OHSLR30 / type: rna / ID: 2 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 20.549197 KDa |
| Sequence | String: GGAUCGAUCG AUCGAUCGGC AUCGAUCGGC UUCGGCCGAU CGAUGCCGAU CGAUCGAUCG AUCC |
-Macromolecule #3: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 3 / Number of copies: 1 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 1 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #5: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 5 / Number of copies: 1 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)