[English] 日本語
 Yorodumi
Yorodumi- EMDB-27170: Human mitochondrial DNA polymerase gamma ternary complex with GT ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human mitochondrial DNA polymerase gamma ternary complex with GT basepair in editing conformer (local refinement of subunit A and primer/template DNA) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | DNA-binding protein / DNA polymerase / TRANSFERASE-DNA complex / TRANSFERASE | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
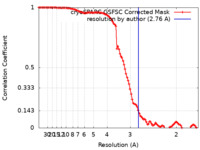
| Method | single particle reconstruction / cryo EM / Resolution: 2.76 Å | |||||||||
 Authors Authors | Park J / Yin YW | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2023 Journal: Nat Struct Mol Biol / Year: 2023Title: Polγ coordinates DNA synthesis and proofreading to ensure mitochondrial genome integrity. Authors: Joon Park / Geoffrey K Herrmann / Patrick G Mitchell / Michael B Sherman / Y Whitney Yin /  Abstract: Accurate replication of mitochondrial DNA (mtDNA) by DNA polymerase γ (Polγ) is essential for maintaining cellular energy supplies, metabolism, and cell cycle control. To illustrate the structural ...Accurate replication of mitochondrial DNA (mtDNA) by DNA polymerase γ (Polγ) is essential for maintaining cellular energy supplies, metabolism, and cell cycle control. To illustrate the structural mechanism for Polγ coordinating polymerase (pol) and exonuclease (exo) activities to ensure rapid and accurate DNA synthesis, we determined four cryo-EM structures of Polγ captured after accurate or erroneous incorporation to a resolution of 2.4-3.0 Å. The structures show that Polγ employs a dual-checkpoint mechanism to sense nucleotide misincorporation and initiate proofreading. The transition from replication to error editing is accompanied by increased dynamics in both DNA and enzyme, in which the polymerase relaxes its processivity and the primer-template DNA unwinds, rotates, and backtracks to shuttle the mismatch-containing primer terminus 32 Å to the exo site for editing. Our structural and functional studies also provide a foundation for analyses of Polγ mutation-induced human diseases and aging. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27170.map.gz emd_27170.map.gz | 59.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27170-v30.xml emd-27170-v30.xml emd-27170.xml emd-27170.xml | 16.5 KB 16.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27170_fsc.xml emd_27170_fsc.xml | 10.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_27170.png emd_27170.png | 141.7 KB | ||
| Masks |  emd_27170_msk_1.map emd_27170_msk_1.map | 125 MB |  Mask map Mask map | |
| Others |  emd_27170_additional_1.map.gz emd_27170_additional_1.map.gz emd_27170_half_map_1.map.gz emd_27170_half_map_1.map.gz emd_27170_half_map_2.map.gz emd_27170_half_map_2.map.gz | 62.7 MB 116.1 MB 116.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27170 http://ftp.pdbj.org/pub/emdb/structures/EMD-27170 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27170 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27170 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_27170.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27170.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_27170_msk_1.map emd_27170_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: CryoSPARC unsharpened map from local refinement
| File | emd_27170_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoSPARC unsharpened map from local refinement | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_27170_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_27170_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human mitochondrial DNA polymerase gamma ternary complex with GT ...
| Entire | Name: Human mitochondrial DNA polymerase gamma ternary complex with GT basepair in editing conformer |
|---|---|
| Components |
|
-Supramolecule #1: Human mitochondrial DNA polymerase gamma ternary complex with GT ...
| Supramolecule | Name: Human mitochondrial DNA polymerase gamma ternary complex with GT basepair in editing conformer type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 44.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 105000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)