[English] 日本語
 Yorodumi
Yorodumi- EMDB-27112: S728-1157 IgG in complex with SARS-CoV-2-6P-Mut7 Spike protein (g... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | S728-1157 IgG in complex with SARS-CoV-2-6P-Mut7 Spike protein (global refinement) | |||||||||
 Map data Map data | sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | SARS-CoV-2 / cross-neutralizing antibody / neutralizing mAb / variants of concern / VIRAL PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
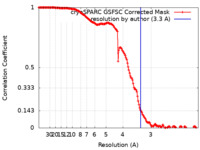
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Ozorowski G / Torres JL / Turner HL / Ward AB | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: J Clin Invest / Year: 2023 Journal: J Clin Invest / Year: 2023Title: Site of vulnerability on SARS-CoV-2 spike induces broadly protective antibody against antigenically distinct Omicron subvariants. Authors: Siriruk Changrob / Peter J Halfmann / Hejun Liu / Jonathan L Torres / Joshua J C McGrath / Gabriel Ozorowski / Lei Li / G Dewey Wilbanks / Makoto Kuroda / Tadashi Maemura / Min Huang / Nai- ...Authors: Siriruk Changrob / Peter J Halfmann / Hejun Liu / Jonathan L Torres / Joshua J C McGrath / Gabriel Ozorowski / Lei Li / G Dewey Wilbanks / Makoto Kuroda / Tadashi Maemura / Min Huang / Nai-Ying Zheng / Hannah L Turner / Steven A Erickson / Yanbin Fu / Atsuhiro Yasuhara / Gagandeep Singh / Brian Monahan / Jacob Mauldin / Komal Srivastava / Viviana Simon / Florian Krammer / D Noah Sather / Andrew B Ward / Ian A Wilson / Yoshihiro Kawaoka / Patrick C Wilson /   Abstract: The rapid evolution of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) Omicron variants has emphasized the need to identify antibodies with broad neutralizing capabilities to inform ...The rapid evolution of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) Omicron variants has emphasized the need to identify antibodies with broad neutralizing capabilities to inform future monoclonal therapies and vaccination strategies. Herein, we identified S728-1157, a broadly neutralizing antibody (bnAb) targeting the receptor-binding site (RBS) that was derived from an individual previously infected with WT SARS-CoV-2 prior to the spread of variants of concern (VOCs). S728-1157 demonstrated broad cross-neutralization of all dominant variants, including D614G, Beta, Delta, Kappa, Mu, and Omicron (BA.1/BA.2/BA.2.75/BA.4/BA.5/BL.1/XBB). Furthermore, S728-1157 protected hamsters against in vivo challenges with WT, Delta, and BA.1 viruses. Structural analysis showed that this antibody targets a class 1/RBS-A epitope in the receptor binding domain via multiple hydrophobic and polar interactions with its heavy chain complementarity determining region 3 (CDR-H3), in addition to common motifs in CDR-H1/CDR-H2 of class 1/RBS-A antibodies. Importantly, this epitope was more readily accessible in the open and prefusion state, or in the hexaproline (6P)-stabilized spike constructs, as compared with diproline (2P) constructs. Overall, S728-1157 demonstrates broad therapeutic potential and may inform target-driven vaccine designs against future SARS-CoV-2 variants. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27112.map.gz emd_27112.map.gz | 324.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27112-v30.xml emd-27112-v30.xml emd-27112.xml emd-27112.xml | 17.1 KB 17.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27112_fsc.xml emd_27112_fsc.xml | 14.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_27112.png emd_27112.png | 123.6 KB | ||
| Masks |  emd_27112_msk_1.map emd_27112_msk_1.map | 343 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-27112.cif.gz emd-27112.cif.gz | 4.7 KB | ||
| Others |  emd_27112_half_map_1.map.gz emd_27112_half_map_1.map.gz emd_27112_half_map_2.map.gz emd_27112_half_map_2.map.gz | 318.8 MB 318.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27112 http://ftp.pdbj.org/pub/emdb/structures/EMD-27112 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27112 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27112 | HTTPS FTP |
-Validation report
| Summary document |  emd_27112_validation.pdf.gz emd_27112_validation.pdf.gz | 933.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_27112_full_validation.pdf.gz emd_27112_full_validation.pdf.gz | 933.2 KB | Display | |
| Data in XML |  emd_27112_validation.xml.gz emd_27112_validation.xml.gz | 24.1 KB | Display | |
| Data in CIF |  emd_27112_validation.cif.gz emd_27112_validation.cif.gz | 31.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27112 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27112 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27112 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27112 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_27112.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27112.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.045 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_27112_msk_1.map emd_27112_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map B
| File | emd_27112_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map A
| File | emd_27112_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : S728-1157 IgG in complex with SARS-CoV-2-6P-Mut7 S protein
| Entire | Name: S728-1157 IgG in complex with SARS-CoV-2-6P-Mut7 S protein |
|---|---|
| Components |
|
-Supramolecule #1: S728-1157 IgG in complex with SARS-CoV-2-6P-Mut7 S protein
| Supramolecule | Name: S728-1157 IgG in complex with SARS-CoV-2-6P-Mut7 S protein type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.7 mg/mL | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: Detergent (LMNG) added shortly before vitrification | ||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE | ||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number real images: 1718 / Average exposure time: 9.0 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)