+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2648 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | stem cell factor-induced intact KIT dimers | |||||||||
 Map data Map data | Reconstruction of intact, kinase-dead KIT-SCF complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | KIT / c-Kit / SCF / stem cell factor / transmembrane / receptor tyrosine kinase / RTK | |||||||||
| Function / homology | :  Function and homology information Function and homology information | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 28.8 Å | |||||||||
 Authors Authors | Opatowsky Y / Bleichert F / Unger VM / Schlessinger J | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2014 Journal: Proc Natl Acad Sci U S A / Year: 2014Title: Structure, domain organization, and different conformational states of stem cell factor-induced intact KIT dimers. Authors: Yarden Opatowsky / Irit Lax / Francisco Tomé / Franziska Bleichert / Vinzenz M Unger / Joseph Schlessinger /  Abstract: Using electron microscopy and fitting of crystal structures, we present the 3D reconstruction of ligand-induced dimers of intact receptor tyrosine kinase, KIT. We observe that KIT protomers form ...Using electron microscopy and fitting of crystal structures, we present the 3D reconstruction of ligand-induced dimers of intact receptor tyrosine kinase, KIT. We observe that KIT protomers form close contacts throughout the entire structure of ligand-bound receptor dimers, and that the dimeric receptors adopt multiple, defined conformational states. Interestingly, the homotypic interactions in the membrane proximal Ig-like domain of the extracellular region differ from those observed in the crystal structure of the unconstrained extracellular regions. We observe two prevalent conformations in which the tyrosine kinase domains interact asymmetrically. The asymmetric arrangement of the cytoplasmic regions may represent snapshots of molecular interactions occurring during trans autophosphorylation. Moreover, the asymmetric arrangements may facilitate specific intermolecular interactions necessary for trans phosphorylation of different KIT autophosphorylation sites that are required for stimulation of kinase activity and recruitment of signaling proteins by activated KIT. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2648.map.gz emd_2648.map.gz | 971.5 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2648-v30.xml emd-2648-v30.xml emd-2648.xml emd-2648.xml | 11 KB 11 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_2648.png emd_2648.png | 501.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2648 http://ftp.pdbj.org/pub/emdb/structures/EMD-2648 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2648 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2648 | HTTPS FTP |
-Validation report
| Summary document |  emd_2648_validation.pdf.gz emd_2648_validation.pdf.gz | 198.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2648_full_validation.pdf.gz emd_2648_full_validation.pdf.gz | 197.8 KB | Display | |
| Data in XML |  emd_2648_validation.xml.gz emd_2648_validation.xml.gz | 4.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2648 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2648 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2648 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2648 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2648.map.gz / Format: CCP4 / Size: 1001 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2648.map.gz / Format: CCP4 / Size: 1001 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of intact, kinase-dead KIT-SCF complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 5.5 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
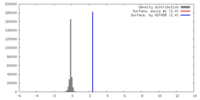
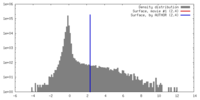
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Intact human KIT dimer in complex with stem cell factor (SCF)dimer
| Entire | Name: Intact human KIT dimer in complex with stem cell factor (SCF)dimer |
|---|---|
| Components |
|
-Supramolecule #1000: Intact human KIT dimer in complex with stem cell factor (SCF)dimer
| Supramolecule | Name: Intact human KIT dimer in complex with stem cell factor (SCF)dimer type: sample / ID: 1000 / Oligomeric state: one dimer of SCF binds to KIT dimer / Number unique components: 2 |
|---|---|
| Molecular weight | Theoretical: 290 KDa / Method: SDS-PAGE |
-Macromolecule #1: stem cell growth factor
| Macromolecule | Name: stem cell growth factor / type: protein_or_peptide / ID: 1 / Name.synonym: SCF / Number of copies: 2 / Oligomeric state: dimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human / Location in cell: Plasma membrane Homo sapiens (human) / synonym: Human / Location in cell: Plasma membrane |
| Molecular weight | Theoretical: 15 KDa |
| Recombinant expression | Organism:  |
| Sequence | UniProtKB: UNIPROTKB: P21583 |
-Macromolecule #2: KIT
| Macromolecule | Name: KIT / type: protein_or_peptide / ID: 2 / Oligomeric state: dimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human / Location in cell: Plasma membrane Homo sapiens (human) / synonym: Human / Location in cell: Plasma membrane |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.05 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: 200mM NaCl, 20mM HEPES,15% glycerol, 0.05% DDM |
| Staining | Type: NEGATIVE Details: grids with adsorbed protein were floated on 3 drops of 1% uranyl formate for 30 seconds each |
| Grid | Details: 400 mesh copper grid with continuous thin carbon support over holy carbon film |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI 12 |
|---|---|
| Date | Nov 27, 2008 |
| Image recording | Category: CCD / Film or detector model: OTHER / Number real images: 400 / Average electron dose: 20 e/Å2 |
| Tilt angle min | 0 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal magnification: 30000 |
| Sample stage | Specimen holder model: OTHER / Tilt angle max: 55 |
- Image processing
Image processing
| Details | Particles of tilt pairs (at 0 and 55 degrees)were manually selected using WEB. Initial reference-free alignment and classification of the untilted views was performed using IMAGIC-5. A representative class average was used for reference-based alignment, followed by cycles of multireference alignment and classification. Three out of 8 classes were used in random conical tilt reconstruction using SPIDER. |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 28.8 Å / Resolution method: OTHER / Software - Name: SPIDER / Number images used: 1900 |
| Final two d classification | Number classes: 3 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name:  Chimera Chimera |
| Details | The domains were separately fitted by manual docking using Chimera |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
-Atomic model buiding 2
| Initial model | PDB ID: |
|---|---|
| Software | Name:  Chimera Chimera |
| Details | The domains were separately fitted by manual docking using Chimera |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)