[English] 日本語
 Yorodumi
Yorodumi- EMDB-26443: Structure of the DU422 SOSIP.664 trimer in complex with neutraliz... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the DU422 SOSIP.664 trimer in complex with neutralizing antibody Fab fragments 10-1074 and BG24 | ||||||||||||
 Map data Map data | Sharpened map | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | broadly neutralizing antibody / bNAb / HIV-1 / CD4 binding site / VH1-2 / VRC01-class / antiviral protein / VIRAL PROTEIN-IMMUNE SYSTEM complex | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell ...symbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / host cell plasma membrane / virion membrane / structural molecule activity / membrane Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /   Human immunodeficiency virus 1 Human immunodeficiency virus 1 | ||||||||||||
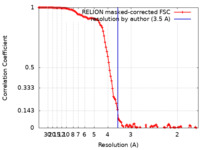
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | ||||||||||||
 Authors Authors | Barnes CO / Bjorkman PJ | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2022 Journal: Sci Adv / Year: 2022Title: A naturally arising broad and potent CD4-binding site antibody with low somatic mutation. Authors: Christopher O Barnes / Till Schoofs / Priyanthi N P Gnanapragasam / Jovana Golijanin / Kathryn E Huey-Tubman / Henning Gruell / Philipp Schommers / Nina Suh-Toma / Yu Erica Lee / Julio C ...Authors: Christopher O Barnes / Till Schoofs / Priyanthi N P Gnanapragasam / Jovana Golijanin / Kathryn E Huey-Tubman / Henning Gruell / Philipp Schommers / Nina Suh-Toma / Yu Erica Lee / Julio C Cetrulo Lorenzi / Alicja Piechocka-Trocha / Johannes F Scheid / Anthony P West / Bruce D Walker / Michael S Seaman / Florian Klein / Michel C Nussenzweig / Pamela J Bjorkman /   Abstract: The induction of broadly neutralizing antibodies (bNAbs) is a potential strategy for a vaccine against HIV-1. However, most bNAbs exhibit features such as unusually high somatic hypermutation, ...The induction of broadly neutralizing antibodies (bNAbs) is a potential strategy for a vaccine against HIV-1. However, most bNAbs exhibit features such as unusually high somatic hypermutation, including insertions and deletions, which make their induction challenging. VRC01-class bNAbs not only exhibit extraordinary breadth and potency but also rank among the most highly somatically mutated bNAbs. Here, we describe a VRC01-class antibody isolated from a viremic controller, BG24, that is much less mutated than most relatives of its class while achieving comparable breadth and potency. A 3.8-Å x-ray crystal structure of a BG24-BG505 Env trimer complex revealed conserved contacts at the gp120 interface characteristic of the VRC01-class Abs, despite lacking common CDR3 sequence motifs. The existence of moderately mutated CD4-binding site (CD4bs) bNAbs such as BG24 provides a simpler blueprint for CD4bs antibody induction by a vaccine, raising the prospect that such an induction might be feasible with a germline-targeting approach. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26443.map.gz emd_26443.map.gz | 15.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26443-v30.xml emd-26443-v30.xml emd-26443.xml emd-26443.xml | 25.4 KB 25.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26443_fsc.xml emd_26443_fsc.xml | 12 KB | Display |  FSC data file FSC data file |
| Images |  emd_26443.png emd_26443.png | 61 KB | ||
| Filedesc metadata |  emd-26443.cif.gz emd-26443.cif.gz | 7.7 KB | ||
| Others |  emd_26443_half_map_1.map.gz emd_26443_half_map_1.map.gz emd_26443_half_map_2.map.gz emd_26443_half_map_2.map.gz | 114.2 MB 114.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26443 http://ftp.pdbj.org/pub/emdb/structures/EMD-26443 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26443 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26443 | HTTPS FTP |
-Related structure data
| Related structure data |  7ucgMC  7uceC  7ucfC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26443.map.gz / Format: CCP4 / Size: 144.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26443.map.gz / Format: CCP4 / Size: 144.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.869 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_26443_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
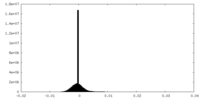
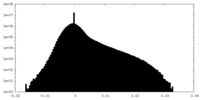
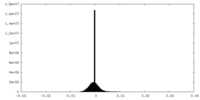
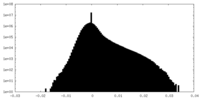
| Density Histograms |
-Half map: #1
| File | emd_26443_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of DU422 SOSIP trimer bound to 10-1074 and BG24 Fab fragments
| Entire | Name: Complex of DU422 SOSIP trimer bound to 10-1074 and BG24 Fab fragments |
|---|---|
| Components |
|
-Supramolecule #1: Complex of DU422 SOSIP trimer bound to 10-1074 and BG24 Fab fragments
| Supramolecule | Name: Complex of DU422 SOSIP trimer bound to 10-1074 and BG24 Fab fragments type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 650 KDa |
-Macromolecule #1: Envelope glycoprotein gp41
| Macromolecule | Name: Envelope glycoprotein gp41 / type: protein_or_peptide / ID: 1 / Details: Env mimic DU422 SOSIP.664 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 17.104527 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AVGLGAVLLG FLGAAGSTMG AASMTLTVQA RQLLSGIVQQ QSNLLRAPEA QQHLLQLTVW GIKQLQTRVL AIERYLKDQQ LLGLWGCSG KLICCTAVPW NSSWSNKSLG DIWDNMTWMQ WDREISNYTN TIFRLLEESQ NQQEKNEKDL LALD UniProtKB: Envelope glycoprotein gp160 |
-Macromolecule #2: BG24 CDRH2-v2 Fab heavy chain
| Macromolecule | Name: BG24 CDRH2-v2 Fab heavy chain / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 25.49859 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLVQSRAE VKKPGASVKV SCEASGYNFV DHYIHWVRQA PGQRPQWVGW MNPQWGQVNY ARTFQGRVTM TRDTSIDTAY MQLNRLTSG DTAVYYCATQ VKLDSSAGYP FDIWGQGTMV TVSSASTKGP SVFPLAPSSK STSGGTAALG CLVKDYFPEP V TVSWNSGA ...String: QVQLVQSRAE VKKPGASVKV SCEASGYNFV DHYIHWVRQA PGQRPQWVGW MNPQWGQVNY ARTFQGRVTM TRDTSIDTAY MQLNRLTSG DTAVYYCATQ VKLDSSAGYP FDIWGQGTMV TVSSASTKGP SVFPLAPSSK STSGGTAALG CLVKDYFPEP V TVSWNSGA LTSGVHTFPA VLQSSGLYSL SSVVTVPSSS LGTQTYICNV NHKPSNTKVD KKVEPKSCDK HHHHHH |
-Macromolecule #3: BG24 light chain
| Macromolecule | Name: BG24 light chain / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 21.925209 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QSALTQPRSV SGSPGQSVNI SCTGAYSGLG WYQQHPGRAP KLIIYEVNRR PSGVSDRFSG SKSGNTASLT ISGLRTEDEA DYFCSAFEY FGEGTKLTVL SQPKAAPSVT LFPPSSEELQ ANKATLVCLI SDFYPGAVTV AWKADSSPVK AGVETTTPSK Q SNNKYAAS ...String: QSALTQPRSV SGSPGQSVNI SCTGAYSGLG WYQQHPGRAP KLIIYEVNRR PSGVSDRFSG SKSGNTASLT ISGLRTEDEA DYFCSAFEY FGEGTKLTVL SQPKAAPSVT LFPPSSEELQ ANKATLVCLI SDFYPGAVTV AWKADSSPVK AGVETTTPSK Q SNNKYAAS SYLSLTPEQW KSHRSYSCQV THEGSTVEKT VAPTECS |
-Macromolecule #4: Envelope glycoprotein gp160
| Macromolecule | Name: Envelope glycoprotein gp160 / type: protein_or_peptide / ID: 4 / Details: Env mimic DU422 SOSIP.664 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 56.99998 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDAMKRGLCC VLLLCGAVFV SPSQEIHARF RRGAENLDLW VTVYYGVPVW KEAKTTLFCA SDAKAYDKEV RNVWATHACV PTDPNPQEI VLENVTENFN MWKNDMVDQM HEDIISLWDQ SLKPCVKLTP LCVTLNCKNV NISANANATA TLNSSMNGEI K NCSFNTTT ...String: MDAMKRGLCC VLLLCGAVFV SPSQEIHARF RRGAENLDLW VTVYYGVPVW KEAKTTLFCA SDAKAYDKEV RNVWATHACV PTDPNPQEI VLENVTENFN MWKNDMVDQM HEDIISLWDQ SLKPCVKLTP LCVTLNCKNV NISANANATA TLNSSMNGEI K NCSFNTTT ELRDKKQKVY ALFYKPDVVP LNGGEHNETG EYILINCNSS TITQACPKVS FDPIPIHYCA PAGYAILKCN NK TFNGTGP CNNVSTVQCT HGIKPVVSTQ LLLNGSLAEE EIIVRSENLT NNIKTIIVHL NKSVEINCTR PNNNTRKSVR IGP GQWFYA TGEIIGDIRE AHCNISRETW NSTLIQVKEK LREHYNKTIK FEPSSGGDLE VTTHSFNCRG EFFYCNTTKL FNET KLFNE SEYVDNKTII LPCRIKQIIN MWQEVGRAMY APPIEGNITC KSNITGLLLT WDGGENSTEG VFRPGGGNMK DNWRS ELYK YKVVEIKPLG VAPTKCKRKV VGR UniProtKB: Envelope glycoprotein gp160 |
-Macromolecule #5: 10-1074 Fab heavy chain
| Macromolecule | Name: 10-1074 Fab heavy chain / type: protein_or_peptide / ID: 5 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 26.389465 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLQESGPG LVKPSETLSV TCSVSGDSMN NYYWTWIRQS PGKGLEWIGY ISDRESATYN PSLNSRVVIS RDTSKNQLSL KLNSVTPAD TAVYYCATAR RGQRIYGVVS FGEFFYYYSM DVWGKGTTVT VSSASTKGPS VFPLAPSSKS TSGGTAALGC L VKDYFPEP ...String: QVQLQESGPG LVKPSETLSV TCSVSGDSMN NYYWTWIRQS PGKGLEWIGY ISDRESATYN PSLNSRVVIS RDTSKNQLSL KLNSVTPAD TAVYYCATAR RGQRIYGVVS FGEFFYYYSM DVWGKGTTVT VSSASTKGPS VFPLAPSSKS TSGGTAALGC L VKDYFPEP VTVSWNSGAL TSGVHTFPAV LQSSGLYSLS SVVTVPSSSL GTQTYICNVN HKPSNTKVDK RVEPKSCDKH HH HHH |
-Macromolecule #6: 10-1074 light chain
| Macromolecule | Name: 10-1074 light chain / type: protein_or_peptide / ID: 6 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.708293 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: SYVRPLSVAL GETARISCGR QALGSRAVQW YQHRPGQAPI LLIYNNQDRP SGIPERFSGT PDINFGTRAT LTISGVEAGD EADYYCHMW DSRSGFSWSF GGATRLTVLG QPKAAPSVFI FPPSDEQLKS GTASVVCLLN NFYPREAKVQ WKVDNALQSG N SQESVTEQ ...String: SYVRPLSVAL GETARISCGR QALGSRAVQW YQHRPGQAPI LLIYNNQDRP SGIPERFSGT PDINFGTRAT LTISGVEAGD EADYYCHMW DSRSGFSWSF GGATRLTVLG QPKAAPSVFI FPPSDEQLKS GTASVVCLLN NFYPREAKVQ WKVDNALQSG N SQESVTEQ DSKDSTYSLS STLTLSKADY EKHKVYACEV THQGLSSPVT KSFNRGEC |
-Macromolecule #10: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 10 / Number of copies: 21 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.5 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 1989 / Average exposure time: 3.5 sec. / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: Correlation coefficient |
| Output model |  PDB-7ucg: |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)