[English] 日本語
 Yorodumi
Yorodumi- EMDB-26275: TMEM106B(120-254) singlet amyloid fibril from frontotemporal loba... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | TMEM106B(120-254) singlet amyloid fibril from frontotemporal lobar degeneration with TDP-43 pathology (FTLD-TDP) type A (case 2) | |||||||||
 Map data Map data | TMEM106B(120-254) singlet amyloid fibril from frontotemporal lobar degeneration with TDP-43 pathology (FTLD-TDP) type A (case 2). | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TMEM106B / Amyloid fibril / PROTEIN FIBRIL | |||||||||
| Function / homology |  Function and homology information Function and homology informationlysosomal protein catabolic process / lysosomal lumen acidification / lysosome localization / regulation of lysosome organization / positive regulation of dendrite development / lysosomal transport / dendrite morphogenesis / lysosome organization / neuron cellular homeostasis / late endosome membrane ...lysosomal protein catabolic process / lysosomal lumen acidification / lysosome localization / regulation of lysosome organization / positive regulation of dendrite development / lysosomal transport / dendrite morphogenesis / lysosome organization / neuron cellular homeostasis / late endosome membrane / ATPase binding / lysosome / endosome / lysosomal membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Fitzpatrick AWP / Stowell MHB | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2022 Journal: Cell / Year: 2022Title: Homotypic fibrillization of TMEM106B across diverse neurodegenerative diseases. Authors: Andrew Chang / Xinyu Xiang / Jing Wang / Carolyn Lee / Tamta Arakhamia / Marija Simjanoska / Chi Wang / Yari Carlomagno / Guoan Zhang / Shikhar Dhingra / Manon Thierry / Jolien Perneel / ...Authors: Andrew Chang / Xinyu Xiang / Jing Wang / Carolyn Lee / Tamta Arakhamia / Marija Simjanoska / Chi Wang / Yari Carlomagno / Guoan Zhang / Shikhar Dhingra / Manon Thierry / Jolien Perneel / Bavo Heeman / Lauren M Forgrave / Michael DeTure / Mari L DeMarco / Casey N Cook / Rosa Rademakers / Dennis W Dickson / Leonard Petrucelli / Michael H B Stowell / Ian R A Mackenzie / Anthony W P Fitzpatrick /    Abstract: Misfolding and aggregation of disease-specific proteins, resulting in the formation of filamentous cellular inclusions, is a hallmark of neurodegenerative disease with characteristic filament ...Misfolding and aggregation of disease-specific proteins, resulting in the formation of filamentous cellular inclusions, is a hallmark of neurodegenerative disease with characteristic filament structures, or conformers, defining each proteinopathy. Here we show that a previously unsolved amyloid fibril composed of a 135 amino acid C-terminal fragment of TMEM106B is a common finding in distinct human neurodegenerative diseases, including cases characterized by abnormal aggregation of TDP-43, tau, or α-synuclein protein. A combination of cryoelectron microscopy and mass spectrometry was used to solve the structures of TMEM106B fibrils at a resolution of 2.7 Å from postmortem human brain tissue afflicted with frontotemporal lobar degeneration with TDP-43 pathology (FTLD-TDP, n = 8), progressive supranuclear palsy (PSP, n = 2), or dementia with Lewy bodies (DLB, n = 1). The commonality of abundant amyloid fibrils composed of TMEM106B, a lysosomal/endosomal protein, to a broad range of debilitating human disorders indicates a shared fibrillization pathway that may initiate or accelerate neurodegeneration. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26275.map.gz emd_26275.map.gz | 12.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26275-v30.xml emd-26275-v30.xml emd-26275.xml emd-26275.xml | 14.4 KB 14.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_26275.png emd_26275.png | 191.5 KB | ||
| Filedesc metadata |  emd-26275.cif.gz emd-26275.cif.gz | 5.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26275 http://ftp.pdbj.org/pub/emdb/structures/EMD-26275 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26275 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26275 | HTTPS FTP |
-Related structure data
| Related structure data |  7u12MC  7u0zC  7u10C  7u11C  7u13C  7u14C  7u15C  7u16C  7u17C  7u18C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_26275.map.gz / Format: CCP4 / Size: 75.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26275.map.gz / Format: CCP4 / Size: 75.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | TMEM106B(120-254) singlet amyloid fibril from frontotemporal lobar degeneration with TDP-43 pathology (FTLD-TDP) type A (case 2). | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.074 Å | ||||||||||||||||||||||||||||||||||||
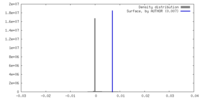
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : TMEM106B(120-254) singlet amyloid fibril from dementia with Lewy ...
| Entire | Name: TMEM106B(120-254) singlet amyloid fibril from dementia with Lewy bodies (DLB) |
|---|---|
| Components |
|
-Supramolecule #1: TMEM106B(120-254) singlet amyloid fibril from dementia with Lewy ...
| Supramolecule | Name: TMEM106B(120-254) singlet amyloid fibril from dementia with Lewy bodies (DLB) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Transmembrane protein 106B
| Macromolecule | Name: Transmembrane protein 106B / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 15.50268 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: SIDVKYIGVK SAYVSYDVQK RTIYLNITNT LNITNNNYYS VEVENITAQV QFSKTVIGKA RLNNITIIGP LDMKQIDYTV PTVIAEEMS YMYDFCTLIS IKVHNIVLMM QVTVTTTYFG HSEQISQERY QYVDCG UniProtKB: Transmembrane protein 106B |
-Macromolecule #2: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 2 / Number of copies: 12 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.8000000000000003 µm / Nominal defocus min: 1.4000000000000001 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 4.8 Å Applied symmetry - Helical parameters - Δ&Phi: -0.4 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Resolution.type: BY AUTHOR / Resolution: 3.5 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 45000 |
|---|---|
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION |
| Startup model | Type of model: NONE |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller


























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)