+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-25753 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Structure of the yeast clamp loader (Replication Factor C RFC) bound to the open sliding clamp (Proliferating Cell Nuclear Antigen PCNA) | |||||||||||||||
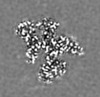
 マップデータ マップデータ | Density modified map (with resolve_cryo_em) | |||||||||||||||
 試料 試料 |
| |||||||||||||||
 キーワード キーワード | sliding clamp / DNA replication / AAA+ / clamp loader / REPLICATION | |||||||||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報Mismatch repair (MMR) directed by MSH2:MSH6 (MutSalpha) / positive regulation of DNA metabolic process / meiotic mismatch repair / Rad17 RFC-like complex / : / Processive synthesis on the lagging strand / Removal of the Flap Intermediate / Elg1 RFC-like complex / DNA replication factor C complex / Ctf18 RFC-like complex ...Mismatch repair (MMR) directed by MSH2:MSH6 (MutSalpha) / positive regulation of DNA metabolic process / meiotic mismatch repair / Rad17 RFC-like complex / : / Processive synthesis on the lagging strand / Removal of the Flap Intermediate / Elg1 RFC-like complex / DNA replication factor C complex / Ctf18 RFC-like complex / : / Polymerase switching / maintenance of DNA trinucleotide repeats / Translesion synthesis by REV1 / : / : / SUMOylation of DNA replication proteins / : / DNA clamp loader activity / DNA replication checkpoint signaling / establishment of mitotic sister chromatid cohesion / : / Activation of ATR in response to replication stress / PCNA complex / lagging strand elongation / DNA damage tolerance / silent mating-type cassette heterochromatin formation / mitotic sister chromatid cohesion / sister chromatid cohesion / error-free translesion synthesis / leading strand elongation / DNA polymerase processivity factor activity / Gap-filling DNA repair synthesis and ligation in TC-NER / Dual incision in TC-NER / subtelomeric heterochromatin formation / mismatch repair / translesion synthesis / DNA damage checkpoint signaling / positive regulation of DNA replication / positive regulation of DNA repair / replication fork / nucleotide-excision repair / DNA-templated DNA replication / mitotic cell cycle / chromosome, telomeric region / cell division / DNA repair / ATP hydrolysis activity / DNA binding / ATP binding / identical protein binding / nucleus / cytosol 類似検索 - 分子機能 | |||||||||||||||
| 生物種 |  | |||||||||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.5 Å | |||||||||||||||
 データ登録者 データ登録者 | Gaubitz C / Demo G / Stone NP / Hayes JA / Liu X / Pajak J / Kelch BA | |||||||||||||||
| 資金援助 |  チェコ, 4件 チェコ, 4件
| |||||||||||||||
 引用 引用 |  ジャーナル: Elife / 年: 2022 ジャーナル: Elife / 年: 2022タイトル: Cryo-EM structures reveal high-resolution mechanism of a DNA polymerase sliding clamp loader. 著者: Christl Gaubitz / Xingchen Liu / Joshua Pajak / Nicholas P Stone / Janelle A Hayes / Gabriel Demo / Brian A Kelch /   要旨: Sliding clamps are ring-shaped protein complexes that are integral to the DNA replication machinery of all life. Sliding clamps are opened and installed onto DNA by clamp loader AAA+ ATPase complexes. ...Sliding clamps are ring-shaped protein complexes that are integral to the DNA replication machinery of all life. Sliding clamps are opened and installed onto DNA by clamp loader AAA+ ATPase complexes. However, how a clamp loader opens and closes the sliding clamp around DNA is still unknown. Here, we describe structures of the clamp loader Replication Factor C (RFC) bound to its cognate sliding clamp Proliferating Cell Nuclear Antigen (PCNA) en route to successful loading. RFC first binds to PCNA in a dynamic, closed conformation that blocks both ATPase activity and DNA binding. RFC then opens the PCNA ring through a large-scale 'crab-claw' expansion of both RFC and PCNA that explains how RFC prefers initial binding of PCNA over DNA. Next, the open RFC:PCNA complex binds DNA and interrogates the primer-template junction using a surprising base-flipping mechanism. Our structures indicate that initial PCNA opening and subsequent closure around DNA do not require ATP hydrolysis, but are driven by binding energy. ATP hydrolysis, which is necessary for RFC release, is triggered by interactions with both PCNA and DNA, explaining RFC's switch-like ATPase activity. Our work reveals how a AAA+ machine undergoes dramatic conformational changes for achieving binding preference and substrate remodeling. | |||||||||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_25753.map.gz emd_25753.map.gz | 8.8 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-25753-v30.xml emd-25753-v30.xml emd-25753.xml emd-25753.xml | 23 KB 23 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_25753_fsc.xml emd_25753_fsc.xml | 8.6 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_25753.png emd_25753.png | 53.4 KB | ||
| Filedesc metadata |  emd-25753.cif.gz emd-25753.cif.gz | 6.4 KB | ||
| その他 |  emd_25753_additional_1.map.gz emd_25753_additional_1.map.gz emd_25753_half_map_1.map.gz emd_25753_half_map_1.map.gz emd_25753_half_map_2.map.gz emd_25753_half_map_2.map.gz | 40.7 MB 40.8 MB 40.8 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25753 http://ftp.pdbj.org/pub/emdb/structures/EMD-25753 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25753 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25753 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_25753.map.gz / 形式: CCP4 / 大きさ: 9.5 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_25753.map.gz / 形式: CCP4 / 大きさ: 9.5 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Density modified map (with resolve_cryo_em) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 これらの図は立方格子座標系で作成されたものです | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
-追加マップ: Unsharpened Map
| ファイル | emd_25753_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Unsharpened Map | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: #1
| ファイル | emd_25753_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: #2
| ファイル | emd_25753_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : Replication Factor C (RFC) bound to the open sliding clamp Prolif...
| 全体 | 名称: Replication Factor C (RFC) bound to the open sliding clamp Proliferating Cell Nuclear Antigen (PCNA) |
|---|---|
| 要素 |
|
-超分子 #1: Replication Factor C (RFC) bound to the open sliding clamp Prolif...
| 超分子 | 名称: Replication Factor C (RFC) bound to the open sliding clamp Proliferating Cell Nuclear Antigen (PCNA) タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: all |
|---|---|
| 由来(天然) | 生物種:  |
| 分子量 | 理論値: 336 KDa |
-分子 #1: PCNA
| 分子 | 名称: PCNA / タイプ: protein_or_peptide / ID: 1 / 光学異性体: LEVO |
|---|---|
| 配列 | 文字列: GPHMASMLEA KFEEASLFKR IIDGFKDCV QLVNFQCKED G IIAQAVDD SRVLLVSLEI GV EAFQEYR CDHPVTLGMD LTS LSKILR CGNNTDTLTL IADN TPDSI ILLFEDTKKD RIAEY SLKL MDIDADFLKI EELQYD STL SLPSSEFSKI VRDLSQL SD ...文字列: GPHMASMLEA KFEEASLFKR IIDGFKDCV QLVNFQCKED G IIAQAVDD SRVLLVSLEI GV EAFQEYR CDHPVTLGMD LTS LSKILR CGNNTDTLTL IADN TPDSI ILLFEDTKKD RIAEY SLKL MDIDADFLKI EELQYD STL SLPSSEFSKI VRDLSQL SD SINIMITKET IKFVADGD I GSGSVIIKPF VDMEHPETS IKLEMDQPVD LTFGAKYLLD IIKGSSLSD RVGIRLSSEA P ALFQFDLK SGFLQFFLAP KF NDEE UniProtKB: DNA sliding clamp PCNA |
-分子 #4: Rfc1
| 分子 | 名称: Rfc1 / タイプ: protein_or_peptide / ID: 4 / 光学異性体: LEVO |
|---|---|
| 配列 | 文字列: MVNISDFFGK NKKSVRSSTS RPTRQVGSSK PEVIDLDTES DQESTNKTPK KMPVSNVIDV SETPEGEKKL PLPAKRKASS PTVKPASSKK TKPSSKSSDS ASNITAQDVL DKIPSLDLSN VHVKENAKFD FKSANSNADP DEIVSEIGSF PEGKPNCLLG LTIVFTGVLP ...文字列: MVNISDFFGK NKKSVRSSTS RPTRQVGSSK PEVIDLDTES DQESTNKTPK KMPVSNVIDV SETPEGEKKL PLPAKRKASS PTVKPASSKK TKPSSKSSDS ASNITAQDVL DKIPSLDLSN VHVKENAKFD FKSANSNADP DEIVSEIGSF PEGKPNCLLG LTIVFTGVLP TLERGASEAL AKRYGARVTK SISSKTSVVV LGDEAGPKKL EKIKQLKIKA IDEEGFKQLI AGMPAEGGDG EAAEKARRKL EEQHNIATKE AELLVKKEEE RSKKLAATRV SGGHLERDNV VREEDKLWTV KYAPTNLQQV CGNKGSVMKL KNWLANWENS KKNSFKHAGK DGSGVFRAAM LYGPPGIGKT TAAHLVAQEL GYDILEQNAS DVRSKTLLNA GVKNALDNMS VVGYFKHNEE AQNLNGKHFV IIMDEVDGMS GGDRGGVGQL AQFCRKTSTP LILICNERNL PKMRPFDRVC LDIQFRRPDA NSIKSRLMTI AIREKFKLDP NVIDRLIQTT RGDIRQVINL LSTISTTTKT INHENINEIS KAWEKNIALK PFDIAHKMLD GQIYSDIGSR NFTLNDKIAL YFDDFDFTPL MIQENYLSTR PSVLKPGQSH LEAVAEAANC ISLGDIVEKK IRSSEQLWSL LPLHAVLSSV YPASKVAGHM AGRINFTAWL GQNSKSAKYY RLLQEIHYHT RLGTSTDKIG LRLDYLPTFR KRLLDPFLKQ GADAISSVIE VMDDYYLTKE DWDSIMEFFV GPDVTTAIIK KIPATVKSGF TRKYNSMTHP VAIYRTGSTI GGGGVGTSTS TPDFEDVVDA DDNPVPADDE ETQDSSTDLK KDKLIKQKAK PTKRKTATSK PGGSKKRKTK A UniProtKB: Replication factor C subunit 1 |
-分子 #5: Rfc2
| 分子 | 名称: Rfc2 / タイプ: protein_or_peptide / ID: 5 / 光学異性体: LEVO |
|---|---|
| 配列 | 文字列: MFEGFGPNKK RKISKLAAEQ SLAQQPWVEK YRPKNLDEVT AQDHAVTVLK KTLKSANLPH MLFYGPPGTG KTSTILALTK ELYGPDLMKS RILELNASDE RGISIVREKV KNFARLTVSK PSKHDLENYP CPPYKIIILD EADSMTADAQ SALRRTMETY SGVTRFCLIC ...文字列: MFEGFGPNKK RKISKLAAEQ SLAQQPWVEK YRPKNLDEVT AQDHAVTVLK KTLKSANLPH MLFYGPPGTG KTSTILALTK ELYGPDLMKS RILELNASDE RGISIVREKV KNFARLTVSK PSKHDLENYP CPPYKIIILD EADSMTADAQ SALRRTMETY SGVTRFCLIC NYVTRIIDPL ASRCSKFRFK ALDASNAIDR LRFISEQENV KCDDGVLERI LDISAGDLRR GITLLQSASK GAQYLGDGKN ITSTQVEELA GVVPHDILIE IVEKVKSGDF DEIKKYVNTF MKSGWSAASV VNQLHEYYIT NDNFDTNFKN QISWLLFTTD SRLNNGTNEH IQLLNLLVKI SQL UniProtKB: Replication factor C subunit 2 |
-分子 #6: Rfc3
| 分子 | 名称: Rfc3 / タイプ: protein_or_peptide / ID: 6 / 光学異性体: LEVO |
|---|---|
| 配列 | 文字列: MSTSTEKRSK ENLPWVEKYR PETLDEVYGQ NEVITTVRKF VDEGKLPHLL FYGPPGTGKT STIVALAREI YGKNYSNMVL ELNASDDRGI DVVRNQIKDF ASTRQIFSKG FKLIILDEAD AMTNAAQNAL RRVIERYTKN TRFCVLANYA HKLTPALLSR CTRFRFQPLP ...文字列: MSTSTEKRSK ENLPWVEKYR PETLDEVYGQ NEVITTVRKF VDEGKLPHLL FYGPPGTGKT STIVALAREI YGKNYSNMVL ELNASDDRGI DVVRNQIKDF ASTRQIFSKG FKLIILDEAD AMTNAAQNAL RRVIERYTKN TRFCVLANYA HKLTPALLSR CTRFRFQPLP QEAIERRIAN VLVHEKLKLS PNAEKALIEL SNGDMRRVLN VLQSCKATLD NPDEDEISDD VIYECCGAPR PSDLKAVLKS ILEDDWGTAH YTLNKVRSAK GLALIDLIEG IVKILEDYEL QNEETRVHLL TKLADIEYSI SKGGNDQIQG SAVIGAIKAS FENETVKANV UniProtKB: Replication factor C subunit 3 |
-分子 #7: Rfc4
| 分子 | 名称: Rfc4 / タイプ: protein_or_peptide / ID: 7 / 光学異性体: LEVO |
|---|---|
| 配列 | 文字列: MSKTLSLQLP WVEKYRPQVL SDIVGNKETI DRLQQIAKDG NMPHMIISGM PGIGKTTSVH CLAHELLGRS YADGVLELNA SDDRGIDVVR NQIKHFAQKK LHLPPGKHKI VILDEADSMT AGAQQALRRT MELYSNSTRF AFACNQSNKI IEPLQSRCAI LRYSKLSDED ...文字列: MSKTLSLQLP WVEKYRPQVL SDIVGNKETI DRLQQIAKDG NMPHMIISGM PGIGKTTSVH CLAHELLGRS YADGVLELNA SDDRGIDVVR NQIKHFAQKK LHLPPGKHKI VILDEADSMT AGAQQALRRT MELYSNSTRF AFACNQSNKI IEPLQSRCAI LRYSKLSDED VLKRLLQIIK LEDVKYTNDG LEAIIFTAEG DMRQAINNLQ STVAGHGLVN ADNVFKIVDS PHPLIVKKML LASNLEDSIQ ILRTDLWKKG YSSIDIVTTS FRVTKNLAQV KESVRLEMIK EIGLTHMRIL EGVGTYLQLA SMLAKIHKLN NKA UniProtKB: Replication factor C subunit 4 |
-分子 #8: Rfc5
| 分子 | 名称: Rfc5 / タイプ: protein_or_peptide / ID: 8 / 光学異性体: LEVO |
|---|---|
| 配列 | 文字列: MSLWVDKYRP KSLNALSHNE ELTNFLKSLS DQPRDLPHLL LYGPNGTGKK TRCMALLESI FGPGVYRLKI DVRQFVTASN RKLELNVVSS PYHLEITPSD MGNNDRIVIQ ELLKEVAQME QVDFQDSKDG LAHRYKCVII NEANSLTKDA QAALRRTMEK YSKNIRLIMV ...文字列: MSLWVDKYRP KSLNALSHNE ELTNFLKSLS DQPRDLPHLL LYGPNGTGKK TRCMALLESI FGPGVYRLKI DVRQFVTASN RKLELNVVSS PYHLEITPSD MGNNDRIVIQ ELLKEVAQME QVDFQDSKDG LAHRYKCVII NEANSLTKDA QAALRRTMEK YSKNIRLIMV CDSMSPIIAP IKSRCLLIRC PAPSDSEIST ILSDVVTNER IQLETKDILK RIAQASNGNL RVSLLMLESM ALNNELALKS SSPIIKPDWI IVIHKLTRKI VKERSVNSLI ECRAVLYDLL AHCIPANIIL KELTFSLLDV ETLNTTNKSS IIEYSSVFDE RLSLGNKAIF HLEGFIAKVM CCLD UniProtKB: Replication factor C subunit 5 |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.5 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | TFS KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K3 (6k x 4k) / 平均電子線量: 40.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD 最大 デフォーカス(公称値): 2.3000000000000003 µm 最小 デフォーカス(公称値): 1.2 µm / 倍率(公称値): 81000 |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー





























 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)