[English] 日本語
 Yorodumi
Yorodumi- EMDB-25621: The Envelope Glycoprotein SIVmac239.K180S SOSIP trimer in complex... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
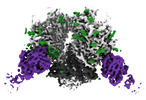
| Title | The Envelope Glycoprotein SIVmac239.K180S SOSIP trimer in complex with 3 copies of the neutralizing antibody K11 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | SIV / Env / vaccine / neutralizing antibody / glycoprotein / VIRAL PROTEIN / VIRAL PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationmembrane fusion involved in viral entry into host cell / host cell endosome membrane / membrane => GO:0016020 / symbiont entry into host cell / viral envelope / virion attachment to host cell / host cell plasma membrane / structural molecule activity / virion membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Simian immunodeficiency virus / Simian immunodeficiency virus /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.47 Å | |||||||||
 Authors Authors | Berndsen ZT / Ward AB | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Molecular insights into antibody-mediated protection against the prototypic simian immunodeficiency virus. Authors: Fangzhu Zhao / Zachary T Berndsen / Nuria Pedreño-Lopez / Alison Burns / Joel D Allen / Shawn Barman / Wen-Hsin Lee / Srirupa Chakraborty / Sandrasegaram Gnanakaran / Leigh M Sewall / ...Authors: Fangzhu Zhao / Zachary T Berndsen / Nuria Pedreño-Lopez / Alison Burns / Joel D Allen / Shawn Barman / Wen-Hsin Lee / Srirupa Chakraborty / Sandrasegaram Gnanakaran / Leigh M Sewall / Gabriel Ozorowski / Oliver Limbo / Ge Song / Peter Yong / Sean Callaghan / Jessica Coppola / Kim L Weisgrau / Jeffrey D Lifson / Rebecca Nedellec / Thomas B Voigt / Fernanda Laurino / Johan Louw / Brandon C Rosen / Michael Ricciardi / Max Crispin / Ronald C Desrosiers / Eva G Rakasz / David I Watkins / Raiees Andrabi / Andrew B Ward / Dennis R Burton / Devin Sok /   Abstract: SIVmac239 infection of macaques is a favored model of human HIV infection. However, the SIVmac239 envelope (Env) trimer structure, glycan occupancy, and the targets and ability of neutralizing ...SIVmac239 infection of macaques is a favored model of human HIV infection. However, the SIVmac239 envelope (Env) trimer structure, glycan occupancy, and the targets and ability of neutralizing antibodies (nAbs) to protect against SIVmac239 remain unknown. Here, we report the isolation of SIVmac239 nAbs that recognize a glycan hole and the V1/V4 loop. A high-resolution structure of a SIVmac239 Env trimer-nAb complex shows many similarities to HIV and SIVcpz Envs, but with distinct V4 features and an extended V1 loop. Moreover, SIVmac239 Env has a higher glycan shield density than HIV Env that may contribute to poor or delayed nAb responses in SIVmac239-infected macaques. Passive transfer of a nAb protects macaques from repeated intravenous SIVmac239 challenge at serum titers comparable to those described for protection of humans against HIV infection. Our results provide structural insights for vaccine design and shed light on antibody-mediated protection in the SIV model. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25621.map.gz emd_25621.map.gz | 97.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25621-v30.xml emd-25621-v30.xml emd-25621.xml emd-25621.xml | 20.9 KB 20.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25621_fsc.xml emd_25621_fsc.xml | 10.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_25621.png emd_25621.png | 127.7 KB | ||
| Masks |  emd_25621_msk_1.map emd_25621_msk_1.map | 103 MB |  Mask map Mask map | |
| Others |  emd_25621_half_map_1.map.gz emd_25621_half_map_1.map.gz emd_25621_half_map_2.map.gz emd_25621_half_map_2.map.gz | 95.4 MB 95.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25621 http://ftp.pdbj.org/pub/emdb/structures/EMD-25621 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25621 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25621 | HTTPS FTP |
-Validation report
| Summary document |  emd_25621_validation.pdf.gz emd_25621_validation.pdf.gz | 833.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_25621_full_validation.pdf.gz emd_25621_full_validation.pdf.gz | 832.9 KB | Display | |
| Data in XML |  emd_25621_validation.xml.gz emd_25621_validation.xml.gz | 18 KB | Display | |
| Data in CIF |  emd_25621_validation.cif.gz emd_25621_validation.cif.gz | 23 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25621 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25621 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25621 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25621 | HTTPS FTP |
-Related structure data
| Related structure data |  7t2pMC  7t4gC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25621.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25621.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.15 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_25621_msk_1.map emd_25621_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
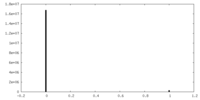
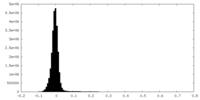
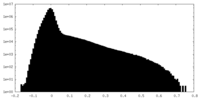
| Density Histograms |
-Half map: #1
| File | emd_25621_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_25621_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : SIVmac239.K180S SOSIP trimer in complex with 3 copies of K11 Fab
| Entire | Name: SIVmac239.K180S SOSIP trimer in complex with 3 copies of K11 Fab |
|---|---|
| Components |
|
-Supramolecule #1: SIVmac239.K180S SOSIP trimer in complex with 3 copies of K11 Fab
| Supramolecule | Name: SIVmac239.K180S SOSIP trimer in complex with 3 copies of K11 Fab type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  Simian immunodeficiency virus Simian immunodeficiency virus |
-Macromolecule #1: Envelope glycoprotein gp120
| Macromolecule | Name: Envelope glycoprotein gp120 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Simian immunodeficiency virus Simian immunodeficiency virus |
| Molecular weight | Theoretical: 60.324441 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGCLGNQLLI AILLLSVYGI YCTLYVTVFY GVPAWRNATI PLFCATKNRD TWGTTQCLPD NGDYSEMALN VTESFDAWNN TVTEQAIED VWQLFETSIK PCVKLSPLCI TMRCNKSETD RWGLTKSITT TASTTSTTAS AKVDMVNETS SCIAQDNCTG L EQEQMISC ...String: MGCLGNQLLI AILLLSVYGI YCTLYVTVFY GVPAWRNATI PLFCATKNRD TWGTTQCLPD NGDYSEMALN VTESFDAWNN TVTEQAIED VWQLFETSIK PCVKLSPLCI TMRCNKSETD RWGLTKSITT TASTTSTTAS AKVDMVNETS SCIAQDNCTG L EQEQMISC KFNMTGLKRD KSKEYNETWY SADLVCEQGN NTGNESRCYM NHCNTSVIQE SCDKHYWDAI RFRYCAPPGY AL LRCNDTN YSGFMPKCSK VVVSSCTRMM ETQTSTWFGF NGTRAENRTY IYWHGRDNRT IISLNKYYNL TMKCRRPGNK TVL PVTIMS GLVFHSQPIN DRPKQAWCWF GGKWKDAIKE VKQTIVKHPR YTGTNNTDKI NLTAPGGGDP EVTFMWTNCR GEFL YCKMN WFLNWVEDRN TANQKPKEQH KRNYVPCHIR QIINTWHKVG KNVYLPPREG DLTCNSTVTS LIANIDWIDG NQTNI TMSA EVAELYRLEL GDYKLVEITP IGLAPTSCKR YTTGGTSRRR RRR |
-Macromolecule #2: Envelope glycoprotein gp41
| Macromolecule | Name: Envelope glycoprotein gp41 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Simian immunodeficiency virus Simian immunodeficiency virus |
| Molecular weight | Theoretical: 18.1446 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GVFVLGFLGF LATAGSAMGA ASLTLTAQSR TLLAGIVQQQ QQLLDVPKRQ QELLRLTVWG TKNLQTRVTA IEKYLKDQAQ LNAWGCAFR QVCCTTVPWP NASLIPKWNN ETWQEWERKV DFLEENITAL LEEAQIQQEK NMYELQKLNG SGHHHHHHHH |
-Macromolecule #3: K11 Fab heavy chain
| Macromolecule | Name: K11 Fab heavy chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 53.29077 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DWTWRILFLV AAATGAHSQV QLQESGPGVV RPSQTLSLTC AVSGDTVSSC CFFWTWIRQP PGKGLEWIGN ISYDNDNTNY NPSLKTRIS ISKDMSKNQF SLKLNSLTAT DTAIYYCARE SPSRGNFCYA YLYGNCPLHF DLWGQGVLVT VSSASTKGPS V FPLAPSSR ...String: DWTWRILFLV AAATGAHSQV QLQESGPGVV RPSQTLSLTC AVSGDTVSSC CFFWTWIRQP PGKGLEWIGN ISYDNDNTNY NPSLKTRIS ISKDMSKNQF SLKLNSLTAT DTAIYYCARE SPSRGNFCYA YLYGNCPLHF DLWGQGVLVT VSSASTKGPS V FPLAPSSR STSESTAALG CLVKDYFPEP VTVSWNSGSL TSGVHTFPAV LQSSGLYSLS SVVTVPSSSL GTQTYVCNVN HK PSNTKVD KRVEIKTCGG GSKPPTCPPC TSPELLGGPS VFLFPPKPKD TLMISRTPEV TCVVVDVSQE DPDVKFNWYV NGA EVHHAQ TKPRETQYNS TYRVVSVLTV THQDWLNGKE YTCKVSNKAL PAPIQKTISK DKGQPREPQV YTLPPSREEL TKNQ VSLTC LVKGFYPSDI VVEWESSGQP ENTYKTTPPV LDSDGSYFLY SKLTVDKSRW QQGNVFSCSV MHEALHNHYT QKSLS LSPG K |
-Macromolecule #4: K11 Fab light chain
| Macromolecule | Name: K11 Fab light chain / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 25.29315 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: ETDTLLLWVL LLWVPGSTGD ILLTQSPSSL SGSVGDRVTI TCRASQGINS YLNWYQQKPG KAPKLLIYFA NRLQSGVPSR FSGSGSGTE FTLTISSLQS EDGATYYCQQ YDTFPTFGPG TKLDIKRTVA APSVFIFPPS EDQVKSGTVS VVCLLNNFYP R EASVKWKV ...String: ETDTLLLWVL LLWVPGSTGD ILLTQSPSSL SGSVGDRVTI TCRASQGINS YLNWYQQKPG KAPKLLIYFA NRLQSGVPSR FSGSGSGTE FTLTISSLQS EDGATYYCQQ YDTFPTFGPG TKLDIKRTVA APSVFIFPPS EDQVKSGTVS VVCLLNNFYP R EASVKWKV DGALKTGNSQ ESVTEQDSKD NTYSLSSTLT LSSTEYQSHK VYACEVTHQG LSSPVTKSFN RGEC |
-Macromolecule #11: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 11 / Number of copies: 13 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Grid | Model: C-flat-1.2/1.3 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z
Z Y
Y X
X