[English] 日本語
 Yorodumi
Yorodumi- EMDB-25572: Pseudomonas phage PaP3 portal protein, delta-C terminal homo-dodecamer -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Pseudomonas phage PaP3 portal protein, delta-C terminal homo-dodecamer | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  Pseudomonas virus PaP3 Pseudomonas virus PaP3 | |||||||||
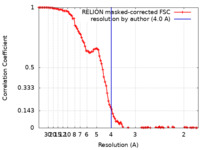
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | |||||||||
 Authors Authors | Hou C-FD / Swanson NA | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2022 Journal: J Mol Biol / Year: 2022Title: Cryo-EM Structure of a Kinetically Trapped Dodecameric Portal Protein from the Pseudomonas-phage PaP3. Authors: Chun-Feng David Hou / Nicholas A Swanson / Fenglin Li / Ruoyu Yang / Ravi K Lokareddy / Gino Cingolani /  Abstract: Portal proteins are dodecameric assemblies that occupy a unique 5-fold vertex of the icosahedral capsid of tailed bacteriophages and herpesviruses. The portal vertex interrupts the icosahedral ...Portal proteins are dodecameric assemblies that occupy a unique 5-fold vertex of the icosahedral capsid of tailed bacteriophages and herpesviruses. The portal vertex interrupts the icosahedral symmetry, and in vivo, its assembly and incorporation in procapsid are controlled by the scaffolding protein. Ectopically expressed portal oligomers are polymorphic in solution, and portal rings built by a different number of subunits have been documented in the literature. In this paper, we describe the cryo-EM structure of the portal protein from the Pseudomonas-phage PaP3, which we determined at 3.4 Å resolution. Structural analysis revealed a dodecamer with helical rather than rotational symmetry, which we hypothesize is kinetically trapped. The helical assembly was stabilized by local mispairing of portal subunits caused by the slippage of crown and barrel helices that move like a lever with respect to the portal body. Removing the C-terminal barrel promoted assembly of undecameric and dodecameric rings with quasi-rotational symmetry, suggesting that the barrel contributes to subunits mispairing. However, ΔC-portal rings were intrinsically asymmetric, with most particles having one open portal subunit interface. Together, these data expand the structural repertoire of viral portal proteins to Pseudomonas-phages and shed light on the unexpected plasticity of the portal protein quaternary structure. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25572.map.gz emd_25572.map.gz | 70.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25572-v30.xml emd-25572-v30.xml emd-25572.xml emd-25572.xml | 15.5 KB 15.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25572_fsc.xml emd_25572_fsc.xml | 10.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_25572.png emd_25572.png | 69.3 KB | ||
| Masks |  emd_25572_msk_1.map emd_25572_msk_1.map | 91.1 MB |  Mask map Mask map | |
| Others |  emd_25572_half_map_1.map.gz emd_25572_half_map_1.map.gz emd_25572_half_map_2.map.gz emd_25572_half_map_2.map.gz | 70.9 MB 70.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25572 http://ftp.pdbj.org/pub/emdb/structures/EMD-25572 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25572 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25572 | HTTPS FTP |
-Validation report
| Summary document |  emd_25572_validation.pdf.gz emd_25572_validation.pdf.gz | 552.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_25572_full_validation.pdf.gz emd_25572_full_validation.pdf.gz | 552.2 KB | Display | |
| Data in XML |  emd_25572_validation.xml.gz emd_25572_validation.xml.gz | 18.1 KB | Display | |
| Data in CIF |  emd_25572_validation.cif.gz emd_25572_validation.cif.gz | 23.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25572 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25572 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25572 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25572 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_25572.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25572.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.91 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_25572_msk_1.map emd_25572_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_25572_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_25572_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Dodecameric complex of phage PaP3 portal, C-terminal truncated mutant
| Entire | Name: Dodecameric complex of phage PaP3 portal, C-terminal truncated mutant |
|---|---|
| Components |
|
-Supramolecule #1: Dodecameric complex of phage PaP3 portal, C-terminal truncated mutant
| Supramolecule | Name: Dodecameric complex of phage PaP3 portal, C-terminal truncated mutant type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Pseudomonas virus PaP3 Pseudomonas virus PaP3 |
| Recombinant expression | Organism:  |
-Macromolecule #1: Pseudomonas phage PaP3 portal protein
| Macromolecule | Name: Pseudomonas phage PaP3 portal protein / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas virus PaP3 Pseudomonas virus PaP3 |
| Sequence | String: MAKRRKIKPM DDEQVLRHLD QLVNDALDFN SSELSKQRSE ALKYYFGEPF GNERPGKSGI VSRDVQETVD WIMPSLMKV FTSGGQVVKY EPDTAEDVEQ AEQETEYVNY LFMRKNEGFK VMFDWFQDTL MMKTGVVKVY V EEVLKPTF ERFSGLSEDM VADILSDPDT ...String: MAKRRKIKPM DDEQVLRHLD QLVNDALDFN SSELSKQRSE ALKYYFGEPF GNERPGKSGI VSRDVQETVD WIMPSLMKV FTSGGQVVKY EPDTAEDVEQ AEQETEYVNY LFMRKNEGFK VMFDWFQDTL MMKTGVVKVY V EEVLKPTF ERFSGLSEDM VADILSDPDT SILAQSVDDD GTYTIKIRKD KKKREIKVLC VKPENFLVDR LA TCIDDAR FLCHREKYTV SDLRLLGVPE DVIEELPYDE YEFSDSQPER LVRDNFDMTG QLQYNSGDDA EAN REVWAS ECYTLLDVDG DGISELRRIL YVGDYIISNE PWDCRPFADL NAYRIAHKFH GMSVYDKIRD IQEI RSVLM RNIMDNIYRT NQGRSVVLDG QVNLEDLLTN EAAGIVRVKS MNSITPLETP QLSGEVYGML DRLEA DRGK RTGITDRTRG LDQNTLHSNQ AAMSVNQLMT AAEQQIDLIA RMFAETGVKR LFQLLHDHAI KYQNQE EVF QLRGKWVAVN PANWRERSDL TVTVGIGNMN KDQQMLHLMR IWEMAQAVVG GGGLGVLVSE QNLYNIL KE VTENAGYKDP DRFWTNPNSP EALQAKAIRE QKEA |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.5 mg/mL | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 278 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 150000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL / Overall B value: 129 / Target criteria: Correlation coefficient |
|---|
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)