[English] 日本語
 Yorodumi
Yorodumi- EMDB-24698: Cryo-EM structure of the HIV-1 restriction factor human SERINC3 -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the HIV-1 restriction factor human SERINC3 | |||||||||
 Map data Map data | WT SERNIC3 masked and sharpened cryo-EM map with CDR of associated Fab. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | flippase / viral restriction / HIV-1 / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationSerine metabolism / L-serine transmembrane transporter activity / phospholipid scramblase activity / positive regulation of endoplasmic reticulum stress-induced intrinsic apoptotic signaling pathway / L-serine biosynthetic process / plasma membrane phospholipid scrambling / antiviral innate immune response / Golgi membrane / perinuclear region of cytoplasm / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.4 Å | |||||||||
 Authors Authors | Purdy MD / Leonhardt SA / Yeager M | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation | Journal: Protein Sci / Year: 2018 Title: UCSF ChimeraX: Meeting modern challenges in visualization and analysis. Authors: Thomas D Goddard / Conrad C Huang / Elaine C Meng / Eric F Pettersen / Gregory S Couch / John H Morris / Thomas E Ferrin /  Abstract: UCSF ChimeraX is next-generation software for the visualization and analysis of molecular structures, density maps, 3D microscopy, and associated data. It addresses challenges in the size, scope, and ...UCSF ChimeraX is next-generation software for the visualization and analysis of molecular structures, density maps, 3D microscopy, and associated data. It addresses challenges in the size, scope, and disparate types of data attendant with cutting-edge experimental methods, while providing advanced options for high-quality rendering (interactive ambient occlusion, reliable molecular surface calculations, etc.) and professional approaches to software design and distribution. This article highlights some specific advances in the areas of visualization and usability, performance, and extensibility. ChimeraX is free for noncommercial use and is available from http://www.rbvi.ucsf.edu/chimerax/ for Windows, Mac, and Linux. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24698.map.gz emd_24698.map.gz | 28.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24698-v30.xml emd-24698-v30.xml emd-24698.xml emd-24698.xml | 22.6 KB 22.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_24698_fsc.xml emd_24698_fsc.xml | 7.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_24698.png emd_24698.png | 84.9 KB | ||
| Masks |  emd_24698_msk_1.map emd_24698_msk_1.map | 30.5 MB |  Mask map Mask map | |
| Others |  emd_24698_half_map_1.map.gz emd_24698_half_map_1.map.gz emd_24698_half_map_2.map.gz emd_24698_half_map_2.map.gz | 23.5 MB 23.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24698 http://ftp.pdbj.org/pub/emdb/structures/EMD-24698 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24698 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24698 | HTTPS FTP |
-Validation report
| Summary document |  emd_24698_validation.pdf.gz emd_24698_validation.pdf.gz | 670.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_24698_full_validation.pdf.gz emd_24698_full_validation.pdf.gz | 670.4 KB | Display | |
| Data in XML |  emd_24698_validation.xml.gz emd_24698_validation.xml.gz | 12.6 KB | Display | |
| Data in CIF |  emd_24698_validation.cif.gz emd_24698_validation.cif.gz | 17.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24698 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24698 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24698 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24698 | HTTPS FTP |
-Related structure data
| Related structure data |  7ru6MC  7rugC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_24698.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24698.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | WT SERNIC3 masked and sharpened cryo-EM map with CDR of associated Fab. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.298 Å | ||||||||||||||||||||||||||||||||||||
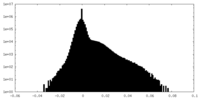
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_24698_msk_1.map emd_24698_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
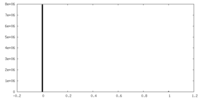
| Density Histograms |
-Half map: WT SERINC3 RELION Refine3D half map used for postprocessing.
| File | emd_24698_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | WT SERINC3 RELION Refine3D half map used for postprocessing. | ||||||||||||
| Projections & Slices |
| ||||||||||||
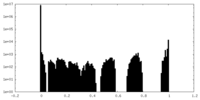
| Density Histograms |
-Half map: WT SERINC3 RELION Refine3D half map used for postprocessing.
| File | emd_24698_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | WT SERINC3 RELION Refine3D half map used for postprocessing. | ||||||||||||
| Projections & Slices |
| ||||||||||||
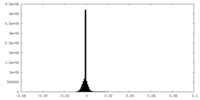
| Density Histograms |
- Sample components
Sample components
-Entire : hSERINC3-SiA
| Entire | Name: hSERINC3-SiA |
|---|---|
| Components |
|
-Supramolecule #1: hSERINC3-SiA
| Supramolecule | Name: hSERINC3-SiA / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all / Details: human WT SERINC3 with synthetic Fab SiA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 55.017 KDa |
-Macromolecule #1: Serine incorporator 3
| Macromolecule | Name: Serine incorporator 3 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 55.05773 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGAVLGVFSL ASWVPCLCSG ASCLLCSCCP NSKNSTVTRL IYAFILLLST VVSYIMQRKE METYLKKIPG FCEGGFKIHE ADINADKDC DVLVGYKAVY RISFAMAIFF FVFSLLMFKV KTSKDLRAAV HNGFWFFKIA ALIGIMVGSF YIPGGYFSSV W FVVGMIGA ...String: MGAVLGVFSL ASWVPCLCSG ASCLLCSCCP NSKNSTVTRL IYAFILLLST VVSYIMQRKE METYLKKIPG FCEGGFKIHE ADINADKDC DVLVGYKAVY RISFAMAIFF FVFSLLMFKV KTSKDLRAAV HNGFWFFKIA ALIGIMVGSF YIPGGYFSSV W FVVGMIGA ALFILIQLVL LVDFAHSWNE SWVNRMEEGN PRLWYAALLS FTSAFYILSI ICVGLLYTYY TKPDGCTENK FF ISINLIL CVVASIISIH PKIQEHQPRS GLLQSSLITL YTMYLTWSAM SNEPDRSCNP NLMSFITRIT APTLAPGNST AVV PTPTPP SKSGSLLDSD NFIGLFVFVL CLLYSSIRTS TNSQVDKLTL SGSDSVILGD TTTSGASDEE DGQPRRAVDN EKEG VQYSY SLFHLMLCLA SLYIMMTLTS WYSPDAKFQS MTSKWPAVWV KISSSWVCLL LYVWTLVAPL VLTSRDFSLE ENLYF QGGS WSHPQFEKAS UniProtKB: Serine incorporator 3 |
-Macromolecule #2: SiA
| Macromolecule | Name: SiA / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 25.621412 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: EISEVQLVES GGGLVQPGGS LRLSCAASGF NFSSSSIHWV RQAPGKGLEW VASISSSSGS TSYADSVKGR FTISADTSKN TAYLQMNSL RAEDTAVYYC ARFYSRYSWY GYSYGWSRAF DYWGQGTLVT VSSASTKGPS VFPLAPSSKS TSGGTAALGC L VKDYFPEP ...String: EISEVQLVES GGGLVQPGGS LRLSCAASGF NFSSSSIHWV RQAPGKGLEW VASISSSSGS TSYADSVKGR FTISADTSKN TAYLQMNSL RAEDTAVYYC ARFYSRYSWY GYSYGWSRAF DYWGQGTLVT VSSASTKGPS VFPLAPSSKS TSGGTAALGC L VKDYFPEP VTVSWNSGAL TSGVHTFPAV LQSSGLYSLS SVVTVPSSSL GTQTYICNVN HKPSNTKVDK KVEPKSCDKT HT C |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: Solutions were made fresh and filtered and degassed. | ||||||||||||
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 2 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 45 sec. / Pretreatment - Atmosphere: AIR | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: blot force 2 blot time 7. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 8710 / Average electron dose: 44.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated defocus max: 1.5 µm / Calibrated defocus min: 0.8 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | An initial hSERINC3 model was generated using AlphaFold v2.1 and the full database excluding PDB:6SP2. Low-confidence loops were removed from the model prior to fitting in the cryoEM map and side chains were retained. |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Overall B value: 147 / Target criteria: Correlation Coefficient |
| Output model |  PDB-7ru6: |
 Movie
Movie Controller
Controller



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)