+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | SthK Y26F Closed State | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cyclic nucleotide-gated ion channel / TRANSPORT PROTEIN | |||||||||
| Biological species |  Spirochaeta thermophila (bacteria) Spirochaeta thermophila (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Gao X / Nimigean C | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Gating intermediates reveal inhibitory role of the voltage sensor in a cyclic nucleotide-modulated ion channel. Authors: Xiaolong Gao / Philipp A M Schmidpeter / Vladimir Berka / Ryan J Durham / Chen Fan / Vasanthi Jayaraman / Crina M Nimigean /   Abstract: Understanding how ion channels gate is important for elucidating their physiological roles and targeting them in pathophysiological states. Here, we used SthK, a cyclic nucleotide-modulated channel ...Understanding how ion channels gate is important for elucidating their physiological roles and targeting them in pathophysiological states. Here, we used SthK, a cyclic nucleotide-modulated channel from Spirochaeta thermophila, to define a ligand-gating trajectory that includes multiple on-pathway intermediates. cAMP is a poor partial agonist for SthK and depolarization increases SthK activity. Tuning the energy landscape by gain-of-function mutations in the voltage sensor domain (VSD) allowed us to capture multiple intermediates along the ligand-activation pathway, highlighting the allosteric linkage between VSD, cyclic nucleotide-binding (CNBD) and pore domains. Small, lateral displacements of the VSD S4 segment were necessary to open the intracellular gate, pointing to an inhibitory VSD at rest. We propose that in wild-type SthK, depolarization leads to such VSD displacements resulting in release of inhibition. In summary, we report conformational transitions along the activation pathway that reveal allosteric couplings between key sites integrating to open the intracellular gate. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24670.map.gz emd_24670.map.gz | 4.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24670-v30.xml emd-24670-v30.xml emd-24670.xml emd-24670.xml | 13.6 KB 13.6 KB | Display Display |  EMDB header EMDB header |
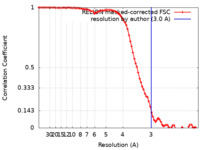
| FSC (resolution estimation) |  emd_24670_fsc.xml emd_24670_fsc.xml | 8.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_24670.png emd_24670.png | 153.3 KB | ||
| Filedesc metadata |  emd-24670.cif.gz emd-24670.cif.gz | 6.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24670 http://ftp.pdbj.org/pub/emdb/structures/EMD-24670 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24670 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24670 | HTTPS FTP |
-Related structure data
| Related structure data |  7rshMC  7rtfC  7rtjC  7ru0C  7ryrC  7rysC M: atomic model generated by this map C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_24670.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24670.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05878 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : SthK
| Entire | Name: SthK |
|---|---|
| Components |
|
-Supramolecule #1: SthK
| Supramolecule | Name: SthK / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 / Details: Y26F mutant of wild type SthK |
|---|---|
| Source (natural) | Organism:  Spirochaeta thermophila (bacteria) Spirochaeta thermophila (bacteria) |
-Macromolecule #1: SthK
| Macromolecule | Name: SthK / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Spirochaeta thermophila (bacteria) Spirochaeta thermophila (bacteria) |
| Molecular weight | Theoretical: 51.102574 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAKDIGINSD PNSSSVDKLM KSSGVSNPTY TLVWKVWILA VTLYFAIRIP LTLVFPSLFS PLLPLDILAS LALIADIPLD LAFESRRTS GRKPTLLAPS RLPDLLAALP LDLLVFALHL PSPLSLLSLV RLLKLISVQR SATRILSYRI NPALLRLLSL V GFILLAAH ...String: MAKDIGINSD PNSSSVDKLM KSSGVSNPTY TLVWKVWILA VTLYFAIRIP LTLVFPSLFS PLLPLDILAS LALIADIPLD LAFESRRTS GRKPTLLAPS RLPDLLAALP LDLLVFALHL PSPLSLLSLV RLLKLISVQR SATRILSYRI NPALLRLLSL V GFILLAAH GIACGWMSLQ PPSENPAGTR YLSAFYWTIT TLTTIGYGDI TPSTPTQTVY TIVIELLGAA MYGLVIGNIA SL VSKLDAA KLLHRERVER VTAFLSYKRI SPELQRRIIE YFDYLWETRR GYEEREVLKE LPHPLRLAVA MEIHGDVIEK VPL FKGAGE EFIRDIILHL EPVIYGPGEY IIRAGEMGSD VYFINRGSVE VLSADEKTRY AILSEGQFFG EMALILRAPR TATV RARAF CDLYRLDKET FDRILSRYPE IAAQIQELAV RRKELESSGL VPRGSVKHHH H |
-Macromolecule #2: ADENOSINE-3',5'-CYCLIC-MONOPHOSPHATE
| Macromolecule | Name: ADENOSINE-3',5'-CYCLIC-MONOPHOSPHATE / type: ligand / ID: 2 / Number of copies: 4 / Formula: CMP |
|---|---|
| Molecular weight | Theoretical: 329.206 Da |
| Chemical component information |  ChemComp-CMP: |
-Macromolecule #3: (1R)-2-{[(S)-{[(2S)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]o...
| Macromolecule | Name: (1R)-2-{[(S)-{[(2S)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]oxy}-1-[(hexadecanoyloxy)methyl]ethyl (9Z)-octadec-9-enoate type: ligand / ID: 3 / Number of copies: 56 / Formula: PGW |
|---|---|
| Molecular weight | Theoretical: 749.007 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 7.5 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7 Component:
| |||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 80 sec. | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV | |||||||||
| Details | Protein in nanodisc of DOPC/POPG/Cardiolipin |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 1.365 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)