[English] 日本語
 Yorodumi
Yorodumi- EMDB-24535: Structure of the human eukaryotic translation initiation factor 2... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-24535 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the human eukaryotic translation initiation factor 2B (eIF2B) in complex with a viral protein NSs | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex / viral protein / integrated stress response / protein translation / TRANSLATION | |||||||||
| Function / homology |  Function and homology information Function and homology informationeukaryotic translation initiation factor 2B complex / Recycling of eIF2:GDP / astrocyte development / astrocyte differentiation / oligodendrocyte development / regulation of translational initiation / cytoplasmic translational initiation / positive regulation of translational initiation / response to glucose / ovarian follicle development ...eukaryotic translation initiation factor 2B complex / Recycling of eIF2:GDP / astrocyte development / astrocyte differentiation / oligodendrocyte development / regulation of translational initiation / cytoplasmic translational initiation / positive regulation of translational initiation / response to glucose / ovarian follicle development / myelination / translation initiation factor binding / translation initiation factor activity / guanyl-nucleotide exchange factor activity / response to endoplasmic reticulum stress / central nervous system development / hippocampus development / translational initiation / response to peptide hormone / regulation of translation / T cell receptor signaling pathway / response to heat / positive regulation of apoptotic process / GTP binding / ATP binding / membrane / identical protein binding / nucleus / plasma membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Sandfly fever Sicilian virus / Sandfly fever Sicilian virus /  Homo sapiens (human) / Homo sapiens (human) /  Sandfly fever sicilian virus Sandfly fever sicilian virus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.6 Å | |||||||||
 Authors Authors | Wang L / Schoof M | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Viral evasion of the integrated stress response through antagonism of eIF2-P binding to eIF2B. Authors: Michael Schoof / Lan Wang / J Zachery Cogan / Rosalie E Lawrence / Morgane Boone / Jennifer Deborah Wuerth / Adam Frost / Peter Walter /   Abstract: Viral infection triggers activation of the integrated stress response (ISR). In response to viral double-stranded RNA (dsRNA), RNA-activated protein kinase (PKR) phosphorylates the translation ...Viral infection triggers activation of the integrated stress response (ISR). In response to viral double-stranded RNA (dsRNA), RNA-activated protein kinase (PKR) phosphorylates the translation initiation factor eIF2, converting it from a translation initiator into a potent translation inhibitor and this restricts the synthesis of viral proteins. Phosphorylated eIF2 (eIF2-P) inhibits translation by binding to eIF2's dedicated, heterodecameric nucleotide exchange factor eIF2B and conformationally inactivating it. We show that the NSs protein of Sandfly Fever Sicilian virus (SFSV) allows the virus to evade the ISR. Mechanistically, NSs tightly binds to eIF2B (K= 30 nM), blocks eIF2-P binding, and rescues eIF2B GEF activity. Cryo-EM structures demonstrate that SFSV NSs and eIF2-P directly compete, with the primary NSs contacts to eIF2Bα mediated by five 'aromatic fingers'. NSs binding preserves eIF2B activity by maintaining eIF2B's conformation in its active A-State. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24535.map.gz emd_24535.map.gz | 230.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24535-v30.xml emd-24535-v30.xml emd-24535.xml emd-24535.xml | 18.3 KB 18.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_24535.png emd_24535.png | 55.9 KB | ||
| Filedesc metadata |  emd-24535.cif.gz emd-24535.cif.gz | 7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24535 http://ftp.pdbj.org/pub/emdb/structures/EMD-24535 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24535 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24535 | HTTPS FTP |
-Related structure data
| Related structure data |  7rloMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_24535.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24535.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.835 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
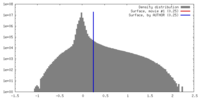
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Eukaryotic translation initiation factor 2B in complex with SFSV NSs
| Entire | Name: Eukaryotic translation initiation factor 2B in complex with SFSV NSs |
|---|---|
| Components |
|
-Supramolecule #1: Eukaryotic translation initiation factor 2B in complex with SFSV NSs
| Supramolecule | Name: Eukaryotic translation initiation factor 2B in complex with SFSV NSs type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Molecular weight | Theoretical: 500 KDa |
-Supramolecule #2: NSs Protein of Sandfly Fever Sicilian Phlebovirus
| Supramolecule | Name: NSs Protein of Sandfly Fever Sicilian Phlebovirus / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #6 |
|---|---|
| Source (natural) | Organism:  Sandfly fever Sicilian virus Sandfly fever Sicilian virus |
-Supramolecule #3: Translation initiation factor eIF-2B subunit
| Supramolecule | Name: Translation initiation factor eIF-2B subunit / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Translation initiation factor eIF-2B subunit epsilon
| Macromolecule | Name: Translation initiation factor eIF-2B subunit epsilon / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 80.466609 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAAPVVAPPG VVVSRANKRS GAGPGGSGGG GARGAEEEPP PPLQAVLVAD SFDRRFFPIS KDQPRVLLPL ANVALIDYTL EFLTATGVQ ETFVFCCWKA AQIKEHLLKS KWCRPTSLNV VRIITSELYR SLGDVLRDVD AKALVRSDFL LVYGDVISNI N ITRALEEH ...String: MAAPVVAPPG VVVSRANKRS GAGPGGSGGG GARGAEEEPP PPLQAVLVAD SFDRRFFPIS KDQPRVLLPL ANVALIDYTL EFLTATGVQ ETFVFCCWKA AQIKEHLLKS KWCRPTSLNV VRIITSELYR SLGDVLRDVD AKALVRSDFL LVYGDVISNI N ITRALEEH RLRRKLEKNV SVMTMIFKES SPSHPTRCHE DNVVVAVDST TNRVLHFQKT QGLRRFAFPL SLFQGSSDGV EV RYDLLDC HISICSPQVA QLFTDNFDYQ TRDDFVRGLL VNEEILGNQI HMHVTAKEYG ARVSNLHMYS AVCADVIRRW VYP LTPEAN FTDSTTQSCT HSRHNIYRGP EVSLGHGSIL EENVLLGSGT VIGSNCFITN SVIGPGCHIG DNVVLDQTYL WQGV RVAAG AQIHQSLLCD NAEVKERVTL KPRSVLTSQV VVGPNITLPE GSVISLHPPD AEEDEDDGEF SDDSGADQEK DKVKM KGYN PAEVGAAGKG YLWKAAGMNM EEEEELQQNL WGLKINMEEE SESESEQSMD SEEPDSRGGS PQMDDIKVFQ NEVLGT LQR GKEENISCDN LVLEINSLKY AYNISLKEVM QVLSHVVLEF PLQQMDSPLD SSRYCALLLP LLKAWSPVFR NYIKRAA DH LEALAAIEDF FLEHEALGIS MAKVLMAFYQ LEILAEETIL SWFSQRDTTD KGQQLRKNQQ LQRFIQWLKE AEEESSED D UniProtKB: Translation initiation factor eIF2B subunit epsilon |
-Macromolecule #2: Translation initiation factor eIF-2B subunit beta
| Macromolecule | Name: Translation initiation factor eIF-2B subunit beta / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 39.039547 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPGSAAKGSE LSERIESFVE TLKRGGGPRS SEEMARETLG LLRQIITDHR WSNAGELMEL IRREGRRMTA AQPSETTVGN MVRRVLKII REEYGRLHGR SDESDQQESL HKLLTSGGLN EDFSFHYAQL QSNIIEAINE LLVELEGTME NIAAQALEHI H SNEVIMTI ...String: MPGSAAKGSE LSERIESFVE TLKRGGGPRS SEEMARETLG LLRQIITDHR WSNAGELMEL IRREGRRMTA AQPSETTVGN MVRRVLKII REEYGRLHGR SDESDQQESL HKLLTSGGLN EDFSFHYAQL QSNIIEAINE LLVELEGTME NIAAQALEHI H SNEVIMTI GFSRTVEAFL KEAARKRKFH VIVAECAPFC QGHEMAVNLS KAGIETTVMT DAAIFAVMSR VNKVIIGTKT IL ANGALRA VTGTHTLALA AKHHSTPLIV CAPMFKLSPQ FPNEEDSFHK FVAPEEVLPF TEGDILEKVS VHCPVFDYVP PEL ITLFIS NIGGNAPSYI YRLMSELYHP DDHVL UniProtKB: Translation initiation factor eIF2B subunit beta |
-Macromolecule #3: Translation initiation factor eIF-2B subunit delta
| Macromolecule | Name: Translation initiation factor eIF-2B subunit delta / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 57.640168 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAAVAVAVRE DSGSGMKAEL PPGPGAVGRE MTKEEKLQLR KEKKQQKKKR KEEKGAEPET GSAVSAAQCQ VGPTRELPES GIQLGTPRE KVPAGRSKAE LRAERRAKQE AERALKQARK GEQGGPPPKA SPSTAGETPS GVKRLPEYPQ VDDLLLRRLV K KPERQQVP ...String: MAAVAVAVRE DSGSGMKAEL PPGPGAVGRE MTKEEKLQLR KEKKQQKKKR KEEKGAEPET GSAVSAAQCQ VGPTRELPES GIQLGTPRE KVPAGRSKAE LRAERRAKQE AERALKQARK GEQGGPPPKA SPSTAGETPS GVKRLPEYPQ VDDLLLRRLV K KPERQQVP TRKDYGSKVS LFSHLPQYSR QNSLTQFMSI PSSVIHPAMV RLGLQYSQGL VSGSNARCIA LLRALQQVIQ DY TTPPNEE LSRDLVNKLK PYMSFLTQCR PLSASMHNAI KFLNKEITSV GSSKREEEAK SELRAAIDRY VQEKIVLAAQ AIS RFAYQK ISNGDVILVY GCSSLVSRIL QEAWTEGRRF RVVVVDSRPW LEGRHTLRSL VHAGVPASYL LIPAASYVLP EVSK VLLGA HALLANGSVM SRVGTAQLAL VARAHNVPVL VCCETYKFCE RVQTDAFVSN ELDDPDDLQC KRGEHVALAN WQNHA SLRL LNLVYDVTPP ELVDLVITEL GMIPCSSVPV VLRVKSSDQ UniProtKB: Translation initiation factor eIF2B subunit delta |
-Macromolecule #4: Translation initiation factor eIF-2B subunit alpha
| Macromolecule | Name: Translation initiation factor eIF-2B subunit alpha / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 33.754148 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDDKELIEYF KSQMKEDPDM ASAVAAIRTL LEFLKRDKGE TIQGLRANLT SAIETLCGVD SSVAVSSGGE LFLRFISLAS LEYSDYSKC KKIMIERGEL FLRRISLSRN KIADLCHTFI KDGATILTHA YSRVVLRVLE AAVAAKKRFS VYVTESQPDL S GKKMAKAL ...String: MDDKELIEYF KSQMKEDPDM ASAVAAIRTL LEFLKRDKGE TIQGLRANLT SAIETLCGVD SSVAVSSGGE LFLRFISLAS LEYSDYSKC KKIMIERGEL FLRRISLSRN KIADLCHTFI KDGATILTHA YSRVVLRVLE AAVAAKKRFS VYVTESQPDL S GKKMAKAL CHLNVPVTVV LDAAVGYIME KADLVIVGAE GVVENGGIIN KIGTNQMAVC AKAQNKPFYV VAESFKFVRL FP LNQQDVP DKFKYKADTL KVAQTGQDLK EEHPWVDYTA PSLITLLFTD LGVLTPSAVS DELIKLYL UniProtKB: Translation initiation factor eIF2B subunit alpha |
-Macromolecule #5: Translation initiation factor eIF-2B subunit gamma
| Macromolecule | Name: Translation initiation factor eIF-2B subunit gamma / type: protein_or_peptide / ID: 5 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 50.30423 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MEFQAVVMAV GGGSRMTDLT SSIPKPLLPV GNKPLIWYPL NLLERVGFEE VIVVTTRDVQ KALCAEFKMK MKPDIVCIPD DADMGTADS LRYIYPKLKT DVLVLSCDLI TDVALHEVVD LFRAYDASLA MLMRKGQDSI EPVPGQKGKK KAVEQRDFIG V DSTGKRLL ...String: MEFQAVVMAV GGGSRMTDLT SSIPKPLLPV GNKPLIWYPL NLLERVGFEE VIVVTTRDVQ KALCAEFKMK MKPDIVCIPD DADMGTADS LRYIYPKLKT DVLVLSCDLI TDVALHEVVD LFRAYDASLA MLMRKGQDSI EPVPGQKGKK KAVEQRDFIG V DSTGKRLL FMANEADLDE ELVIKGSILQ KHPRIRFHTG LVDAHLYCLK KYIVDFLMEN GSITSIRSEL IPYLVRKQFS SA SSQQGQE EKEEDLKKKE LKSLDIYSFI KEANTLNLAP YDACWNACRG DRWEDLSRSQ VRCYVHIMKE GLCSRVSTLG LYM EANRQV PKLLSALCPE EPPVHSSAQI VSKHLVGVDS LIGPETQIGE KSSIKRSVIG SSCLIKDRVT ITNCLLMNSV TVEE GSNIQ GSVICNNAVI EKGADIKDCL IGSGQRIEAK AKRVNEVIVG NDQLMEI UniProtKB: Translation initiation factor eIF2B subunit gamma |
-Macromolecule #6: Non-structural protein NS-S
| Macromolecule | Name: Non-structural protein NS-S / type: protein_or_peptide / ID: 6 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Sandfly fever sicilian virus Sandfly fever sicilian virus |
| Molecular weight | Theoretical: 30.668383 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MNSQYMFDYP AINIDVRCHR LLSSVSYVAY NKFHTHDVST YEHCEIPLEK LRLGFGRRNS LADFYSLGEL PASWGPACYF SSVKPMMYT FQGMASDLSR FDLTSFSRKG LPNVLKALSW PLGIPDCEIF SICSDRFVRG LQTRDQLMSY ILRMGDSHSL D ECIVQAHK ...String: MNSQYMFDYP AINIDVRCHR LLSSVSYVAY NKFHTHDVST YEHCEIPLEK LRLGFGRRNS LADFYSLGEL PASWGPACYF SSVKPMMYT FQGMASDLSR FDLTSFSRKG LPNVLKALSW PLGIPDCEIF SICSDRFVRG LQTRDQLMSY ILRMGDSHSL D ECIVQAHK KILQEARRLG LSDEHYNGYD LFREIGSLVC LRLINAEPFD TASSGEALDV RTVIRSYRAS DPSTGLTEYG NS LWTPIHS HVDENDESSS DSDFGSHHHH HH UniProtKB: Non-structural protein NS-S |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 67.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.6 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 137093 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)